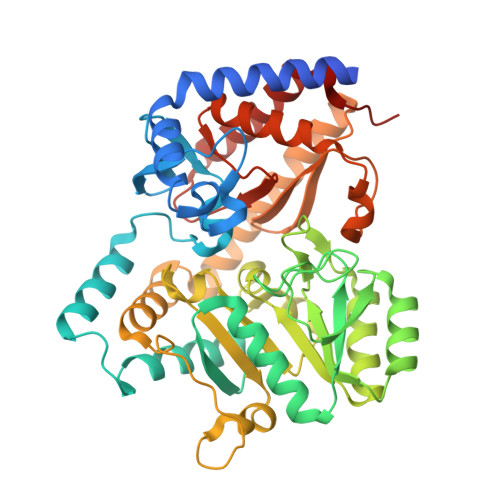
Biochemical Properties and Crystal Structure of a Novel Beta-Phenylalanine Aminotransferase from Variovorax Paradoxus
Crismaru, C.G., Wybenga, G.G., Szymanski, W., Wijma, H.J., Wu, B., Dewildeman, S., Poelarends, G.J., Dijkstra, B.W., Janssen, D.B.(2013) Appl Environ Microbiol 79: 185
- PubMed: 23087034
- DOI: https://doi.org/10.1128/AEM.02525-12
- Primary Citation Related Structures:
4AO9, 4AOA - PubMed Abstract:
By selective enrichment, we isolated a bacterium that can use β-phenylalanine as a sole nitrogen source. It was identified by 16S rRNA gene sequencing as a strain of Variovorax paradoxus. Enzyme assays revealed an aminotransferase activity. Partial genome sequencing and screening of a cosmid DNA library resulted in the identification of a 1,302-bp aminotransferase gene, which encodes a 46,416-Da protein. The gene was cloned and overexpressed in Escherichia coli. The recombinant enzyme was purified and showed a specific activity of 17.5 U mg(-1) for (S)-β-phenylalanine at 30°C and 33 U mg(-1) at the optimum temperature of 55°C. The β-specific aminotransferase exhibits a broad substrate range, accepting ortho-, meta-, and para-substituted β-phenylalanine derivatives as amino donors and 2-oxoglutarate and pyruvate as amino acceptors. The enzyme is highly enantioselective toward (S)-β-phenylalanine (enantioselectivity [E], >100) and derivatives thereof with different substituents on the phenyl ring, allowing the kinetic resolution of various racemic β-amino acids to yield (R)-β-amino acids with >95% enantiomeric excess (ee). The crystal structures of the holoenzyme and of the enzyme in complex with the inhibitor 2-aminooxyacetate revealed structural similarity to the β-phenylalanine aminotransferase from Mesorhizobium sp. strain LUK. The crystal structure was used to rationalize the stereo- and regioselectivity of V. paradoxus aminotransferase and to define a sequence motif with which new aromatic β-amino acid-converting aminotransferases may be identified.
- Department of Biochemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Groningen, The Netherlands.
Organizational Affiliation: