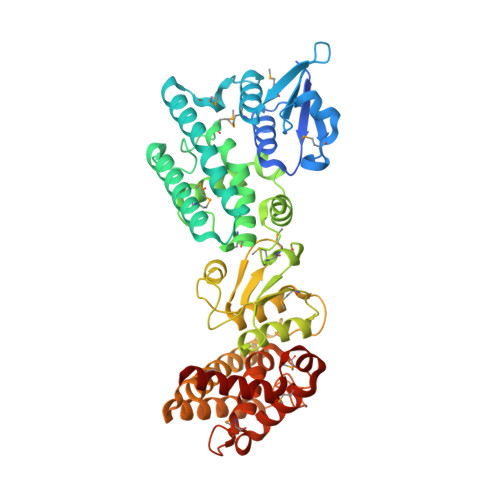
Leishmania Tdr1 Structure, a Unique Trimeric Glutathione Transferase Capable of Deglutathionylation and Antimonial Prodrug Activation.
Fyfe, P.K., Westrop, G.D., Silva, A.M., Coombs, G.H., Hunter, W.N.(2012) Proc Natl Acad Sci U S A 109: 11693
- PubMed: 22753509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1202593109
- Primary Citation Related Structures:
4AGS - PubMed Abstract:
Thiol-dependent reductase I (TDR1), an enzyme found in parasitic Leishmania species and Trypanosoma cruzi, is implicated in deglutathionylation and activation of antimonial prodrugs used to treat leishmaniasis. The 2.3 Å resolution structure of TDR1 reveals a unique trimer of subunits each containing two glutathione-S-transferase (GST) domains. The similarities of individual domains and comparisons with GST classes suggest that TDR1 evolved by gene duplication, diversification, and gene fusion; a combination of events previously unknown in the GST protein superfamily and potentially explaining the distinctive enzyme properties of TDR1. The deglutathionylation activity of TDR1 implies that glutathione itself has regulatory intracellular roles in addition to being a precursor for trypanothione, the major low mass thiol present in trypanosomatids. We propose that activation of antiparasite Sb(V)-drugs is a legacy of the deglutathionylation activity of TDR1 and involves processing glutathione adducts with concomitant reduction of the metalloid to active Sb(III) species.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: