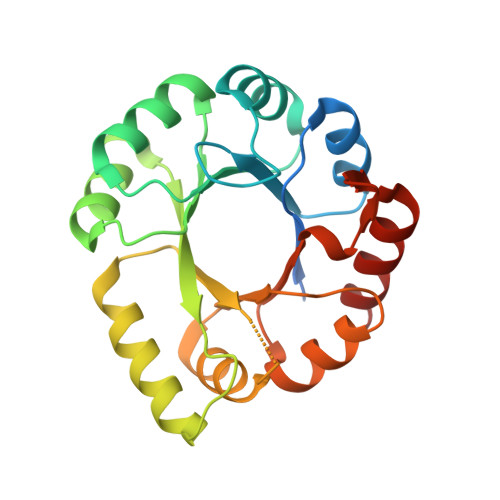
A Monomeric Tim-Barrel Structure from Pyrococcus Furiosus is Optimized for Extreme Temperatures
Repo, H., Oeemig, J.S., Djupsjobacka, J.B., Iwai, H., Heikinheimo, P.(2012) Acta Crystallogr D Biol Crystallogr 68: 1479
- PubMed: 23090397 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912037171
- Primary Citation Related Structures:
4AAJ - PubMed Abstract:
The structure of phosphoribosyl anthranilate isomerase (TrpF) from the hyperthermophilic archaeon Pyrococcus furiosus (PfTrpF) has been determined at 1.75 Å resolution. The PfTrpF structure has a monomeric TIM-barrel fold which differs from the dimeric structures of two other known thermophilic TrpF proteins. A comparison of the PfTrpF structure with the two known bacterial thermophilic TrpF structures and the structure of a related mesophilic protein from Escherichia coli (EcTrpF) is presented. The thermophilic TrpF structures contain a higher proportion of ion pairs and charged residues compared with the mesophilic EcTrpF. These residues contribute to the closure of the central barrel and the stabilization of the barrel and the surrounding α-helices. In the monomeric PfTrpF conserved structural water molecules are mostly absent; instead, the structural waters are replaced by direct side-chain-main-chain interactions. As a consequence of these combined mechanisms, the P. furiosus enzyme is a thermodynamically stable and entropically optimized monomeric TIM-barrel enzyme which defines a good framework for further protein engineering for industrial applications.
- Institute of Biotechnology, University of Helsinki, PO Box 65, FI-00014 Helsinki, Finland.
Organizational Affiliation: