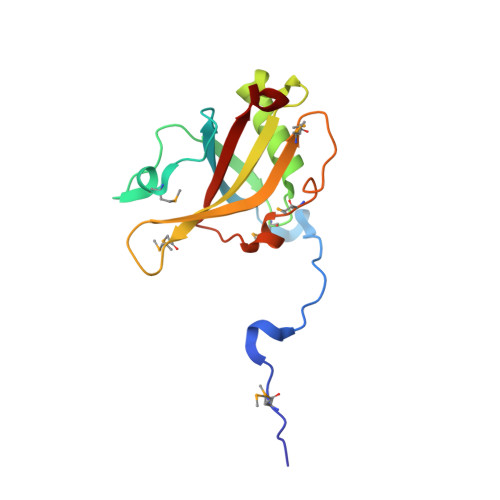
The Structure of the Asap Core Complex Reveals the Existence of a Pinin-Containing Psap Complex
Murachelli, A.G., Ebert, J., Basquin, C., Le Hir, H., Conti, E.(2012) Nat Struct Mol Biol 19: 378
- PubMed: 22388736 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2242
- Primary Citation Related Structures:
4A6Q, 4A8X, 4A90 - PubMed Abstract:
The ASAP complex interacts with the exon-junction complex (EJC), a messenger ribonucleoprotein complex involved in post-transcriptional regulation. The three ASAP subunits (Acinus, RNPS1 and SAP18) have been individually implicated in transcriptional regulation, pre-mRNA splicing and mRNA quality control. To shed light on the basis for and consequences of ASAP's interaction with the EJC, we have determined the 1.9-Å resolution structure of a eukaryotic ASAP core complex. The RNA-recognition motif of RNPS1 binds to a conserved motif of Acinus with a recognition mode similar to that observed in splicing U2AF proteins. The Acinus-RNPS1 platform recruits the ubiquitin-like domain of SAP18, forming a ternary complex that has both RNA- and protein-binding properties. Unexpectedly, our structural analysis identified an Acinus-like motif in Pinin, another EJC-associated splicing factor. We show that Pinin physically interacts with RNPS1 and SAP18, forming an alternative ternary complex, PSAP.
- Max Planck Institute of Biochemistry, Department of Structural Cell Biology, Martinsried, Germany.
Organizational Affiliation: