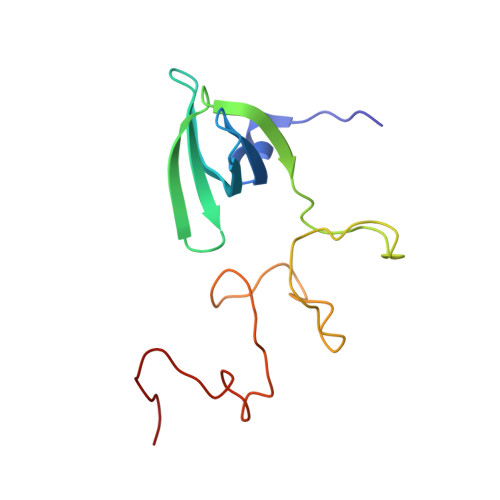
The Structural Basis of Edc3- and Scd6-Mediated Activation of the Dcp1:Dcp2 Mrna Decapping Complex.
Fromm, S.A., Truffault, V., Kamenz, J., Braun, J.E., Hoffmann, N.A., Izaurralde, E., Sprangers, R.(2011) EMBO J 31: 279
- PubMed: 22085934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.408
- Primary Citation Related Structures:
4A53, 4A54 - PubMed Abstract:
The Dcp1:Dcp2 decapping complex catalyses the removal of the mRNA 5' cap structure. Activator proteins, including Edc3 (enhancer of decapping 3), modulate its activity. Here, we solved the structure of the yeast Edc3 LSm domain in complex with a short helical leucine-rich motif (HLM) from Dcp2. The motif interacts with the monomeric Edc3 LSm domain in an unprecedented manner and recognizes a noncanonical binding surface. Based on the structure, we identified additional HLMs in the disordered C-terminal extension of Dcp2 that can interact with Edc3. Moreover, the LSm domain of the Edc3-related protein Scd6 competes with Edc3 for the interaction with these HLMs. We show that both Edc3 and Scd6 stimulate decapping in vitro, presumably by preventing the Dcp1:Dcp2 complex from adopting an inactive conformation. In addition, we show that the C-terminal HLMs in Dcp2 are necessary for the localization of the Dcp1:Dcp2 decapping complex to P-bodies in vivo. Unexpectedly, in contrast to yeast, in metazoans the HLM is found in Dcp1, suggesting that details underlying the regulation of mRNA decapping changed throughout evolution.
- Max Planck Institute for Developmental Biology, Tübingen, Germany.
Organizational Affiliation: