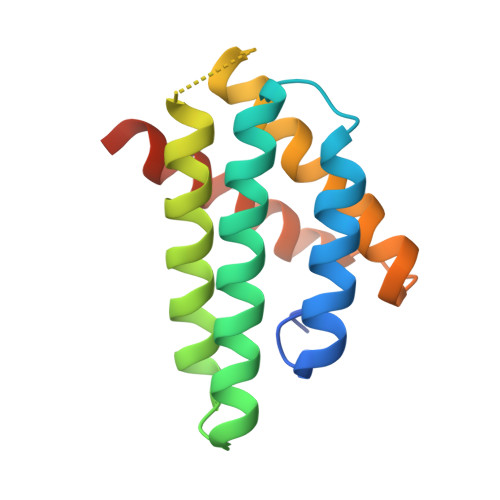
pH-Dependent Dimerization of Spider Silk N-Terminal Domain Requires Relocation of a Wedged Tryptophan Side Chain
Jaudzems, K., Askarieh, G., Landreh, M., Nordling, K., Hedhammar, M., Jornvall, H., Rising, A., Knight, S.D., Johansson, J.(2012) J Mol Biology
- PubMed: 22706024 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.06.004
- Primary Citation Related Structures:
2LPI, 2LPJ, 4FBS - PubMed Abstract:
Formation of spider silk from its constituent proteins-spidroins-involves changes from soluble helical/coil conformations to insoluble β-sheet aggregates. This conversion needs to be regulated to avoid precocious aggregation proximally in the silk gland while still allowing rapid silk assembly in the distal parts. Lowering of pH from about 7 to 6 is apparently important for silk formation. The spidroin N-terminal domain (NT) undergoes stable dimerization and structural changes in this pH region, but the underlying mechanisms are incompletely understood. Here, we determine the NMR and crystal structures of Euprosthenops australis NT mutated in the dimer interface (A72R). Also, the NMR structure of wild-type (wt) E. australis NT at pH7.2 and 300 mM sodium chloride was determined. The wt NT and A72R structures are monomers and virtually identical, but they differ from the subunit structure of dimeric wt NT mainly by having a tryptophan (W10) buried between helix 1 and helix 3, while W10 is surface exposed in the dimer. Wedging of the W10 side chain in monomeric NT tilts helix 3 approximately 5-6Å into a position that is incompatible with that of the observed dimer structure. The structural differences between monomeric and dimeric NT domains explain the tryptophan fluorescence patterns of NT at pH7 and pH6 and indicate that the biological function of NT depends on conversion between the two conformations.
- Department of Physical Organic Chemistry, Latvian Institute of Organic Synthesis, Aizkraukles 21, Riga LV-1006, Latvia.
Organizational Affiliation: