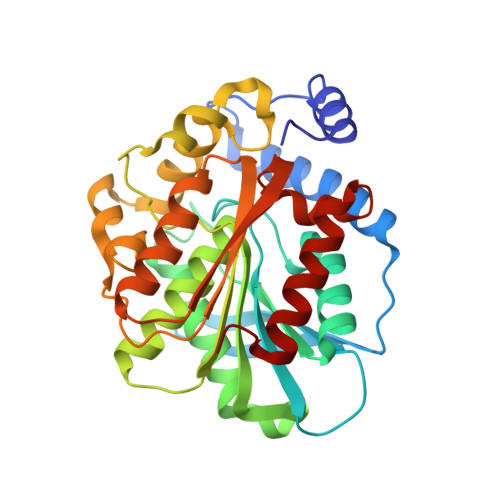
The Crystal Structure of an Esterase from the Hyperthermophilic Microorganism Pyrobaculum Calidifontis Va1 Supports Explanation of its Enantioselectivity.
Palm, G.J., Fernandez-Alvaro, E., Bogdanovic, X., Bartsch, S., Sczodrok, J., Singh, R.K., Boettcher, D., Atomi, H., Bornscheuer, U.T., Hinrichs, W.(2011) Appl Microbiol Biotechnol 91: 1061
- PubMed: 21614503 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-011-3337-9
- Primary Citation Related Structures:
2YH2, 3ZWQ - PubMed Abstract:
The highly thermostable esterase from the hyperthermophilic archaeon Pyrobaculum calidifontis VA1 (PestE) shows high enantioselectivity (E > 100) in the kinetic resolution of racemic chiral carboxylic acids, but little selectivity towards acetates of tertiary alcohols (E = 2-4). To explain these unique properties, its crystal structure has been determined at 2.0 Å resolution. The enzyme is a member of the hormone-sensitive lipase group (group H) of the esterase/lipase superfamily on the basis of the amino acid sequence identity. The PestE structure shows a canonical α/β-hydrolase fold as core domain with a cap structure at the C-terminal end of the β-sheet. A tetramer in the crystal packing is formed of two dimers; the dimeric form is observed in solution. Conserved dimers and even tetramers are found in other group H proteins. The amino acid residues Ser157, His284, and Asp254 form the catalytic triad, which is typically found in α/β-hydrolases. The oxyanion hole is composed of Gly85 and Gly86 within the conserved sequence motif HGGG(M,F,W) (amino acid residues 83-87) and Ala158. With the elucidated structure, experimental results about enantioselectivity towards the two model substrate classes (as exemplified for 3-phenylbutanoic acid ethyl ester and 1,1,1-trifluoro-2-phenylbut-3-yn-2-yl acetate) could be explained by molecular modeling. For both enantiomers of the tertiary alcohol, orientations in two binding pockets were obtained without significant energy differences corresponding to the observed low enantioselectivity due to missing steric repulsions. In contrast, for the carboxylic acid ester, two different orientations with significant energy differences for each enantiomer were found matching the high E values.
- Department of Molecular Structural Biology, Institute of Biochemistry, Greifswald University, Felix-Hausdorff Str. 4, 17487 Greifswald, Germany.
Organizational Affiliation: