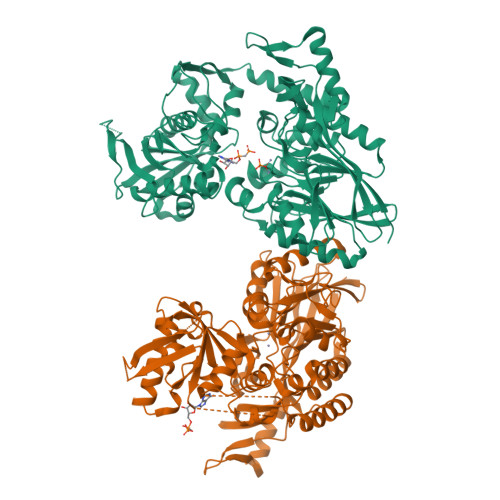
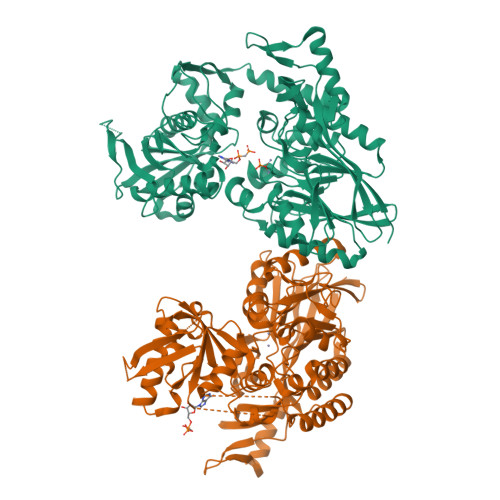
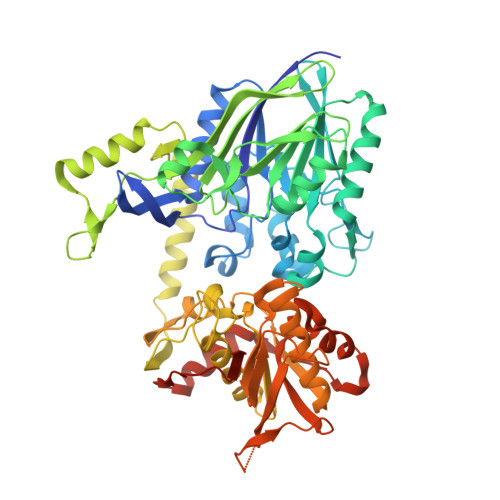
The High-Resolution Crystal Structure of Periplasmic Haemophilus Influenzae Nad Nucleotidase Reveals a Novel Enzymatic Function of Human Cd73 Related to Nad Metabolism.
Garavaglia, S., Bruzzone, S., Cassani, C., Canella, L., Allegrone, G., Sturla, L., Mannino, E., Millo, E., De Flora, A., Rizzi, M.(2012) Biochem J 441: 131
- PubMed: 21933152
- DOI: https://doi.org/10.1042/BJ20111263
- Primary Citation of Related Structures:
3ZTV, 3ZU0 - PubMed Abstract:
Haemophilus influenzae is a major pathogen of the respiratory tract in humans that has developed the capability to exploit host NAD(P) for its nicotinamide dinucleotide requirement. This strategy is organized around a periplasmic enzyme termed NadN (NAD nucleotidase), which plays a central role by degrading NAD into adenosine and NR (nicotinamide riboside), the latter being subsequently internalized by a specific permease. We performed a biochemical and structural investigation on H. influenzae NadN which determined that the enzyme is a Zn2+-dependent 5'-nucleotidase also endowed with NAD(P) pyrophosphatase activity. A 1.3 Å resolution structural analysis revealed a remarkable conformational change that occurs during catalysis between the open and closed forms of the enzyme. NadN showed a broad substrate specificity, recognizing either mono- or di-nucleotide nicotinamides and different adenosine phosphates with a maximal activity on 5'-adenosine monophosphate. Sequence and structural analysis of H. influenzae NadN led us to discover that human CD73 is capable of processing both NAD and NMN, therefore disclosing a possible novel function of human CD73 in systemic NAD metabolism. Our data may prove to be useful for inhibitor design and disclosed unanticipated fascinating evolutionary relationships.
Organizational Affiliation:
Dipartimento di Scienze Chimiche, Alimentari, Farmaceutiche e Farmacologiche, University of Piemonte Orientale Amedeo Avogadro, Via Bovio 6, 28100 Novara, Italy. silvia.garavaglia@pharm.unipmn.it