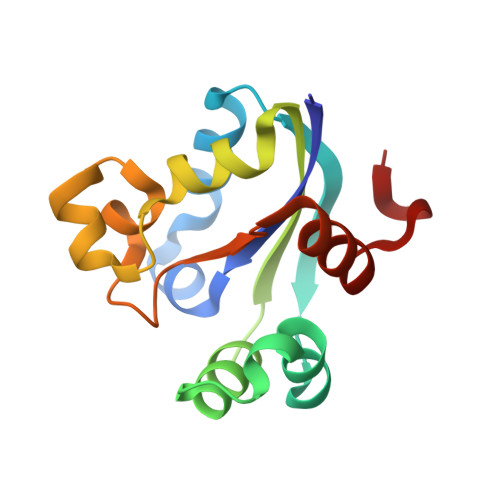
An Intersubunit Disulfide Bridge Stabilizes the Tetrameric Nucleoside Diphosphate Kinase of Aquifex Aeolicus.
Boissier, F., Georgescauld, F., Moynie, L., Dupuy, J.W., Sarger, C., Podar, M., Lascu, I., Giraud, M.F., Dautant, A.(2012) Proteins 80: 1658
- PubMed: 22467275 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24062
- Primary Citation Related Structures:
3ZTO, 3ZTP, 3ZTQ, 3ZTR, 3ZTS - PubMed Abstract:
The nucleoside diphosphate kinase (Ndk) catalyzes the reversible transfer of the γ-phosphate from nucleoside triphosphate to nucleoside diphosphate. Ndks form hexamers or two types of tetramers made of the same building block, namely, the common dimer. The secondary interfaces of the Type I tetramer found in Myxococcus xanthus Ndk and of the Type II found in Escherichia coli Ndk involve the opposite sides of subunits. Up to now, the few available structures of Ndk from thermophiles were hexameric. Here, we determined the X-ray structures of four crystal forms of the Ndk from the hyperthermophilic bacterium Aquifex aeolicus (Aa-Ndk). Aa-Ndk displays numerous features of thermostable proteins and is made of the common dimer but it is a tetramer of Type I. Indeed, the insertion of three residues in a surface-exposed spiral loop, named the Kpn-loop, leads to the formation of a two-turn α-helix that prevents both hexamer and Type II tetramer assembly. Moreover, the side chain of the cysteine at position 133, which is not present in other Ndk sequences, adopts two alternate conformations. Through the secondary interface, each one forms a disulfide bridge with the equivalent Cys133 from the neighboring subunit. This disulfide bridge was progressively broken during X-ray data collection by radiation damage. Such crosslinks counterbalance the weakness of the common-dimer interface. A 40% decrease of the kinase activity at 60°C after reduction and alkylation of the protein corroborates the structural relevance of the disulfide bridge on the tetramer assembly and enzymatic function.
- Université de Bordeaux, Institut de Biochimie et de Génétique Cellulaires, UMR 5095, F-33077 Bordeaux, France.
Organizational Affiliation: