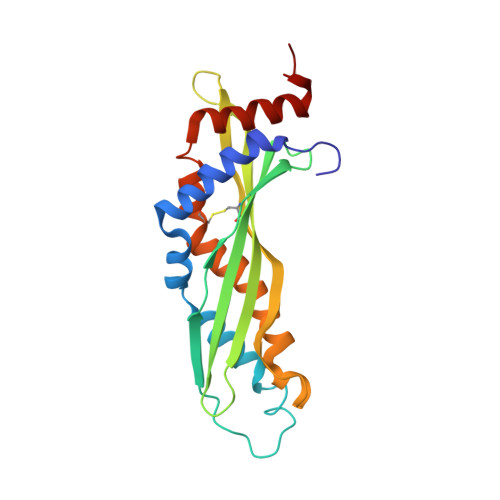
The structure of latherin, a surfactant allergen protein from horse sweat and saliva.
Vance, S.J., McDonald, R.E., Cooper, A., Smith, B.O., Kennedy, M.W.(2013) J R Soc Interface 10: 20130453-20130453
- PubMed: 23782536
- DOI: https://doi.org/10.1098/rsif.2013.0453
- Primary Citation of Related Structures:
3ZPM - PubMed Abstract:
Latherin is a highly surface-active allergen protein found in the sweat and saliva of horses and other equids. Its surfactant activity is intrinsic to the protein in its native form, and is manifest without associated lipids or glycosylation. Latherin probably functions as a wetting agent in evaporative cooling in horses, but it may also assist in mastication of fibrous food as well as inhibition of microbial biofilms. It is a member of the PLUNC family of proteins abundant in the oral cavity and saliva of mammals, one of which has also been shown to be a surfactant and capable of disrupting microbial biofilms. How these proteins work as surfactants while remaining soluble and cell membrane-compatible is not known. Nor have their structures previously been reported. We have used protein nuclear magnetic resonance spectroscopy to determine the conformation and dynamics of latherin in aqueous solution. The protein is a monomer in solution with a slightly curved cylindrical structure exhibiting a 'super-roll' motif comprising a four-stranded anti-parallel β-sheet and two opposing α-helices which twist along the long axis of the cylinder. One end of the molecule has prominent, flexible loops that contain a number of apolar amino acid side chains. This, together with previous biophysical observations, leads us to a plausible mechanism for surfactant activity in which the molecule is first localized to the non-polar interface via these loops, and then unfolds and flattens to expose its hydrophobic interior to the air or non-polar surface. Intrinsically surface-active proteins are relatively rare in nature, and this is the first structure of such a protein from mammals to be reported. Both its conformation and proposed method of action are different from other, non-mammalian surfactant proteins investigated so far.
- School of Chemistry, University of Glasgow, Glasgow G12 8QQ, UK.
Organizational Affiliation: