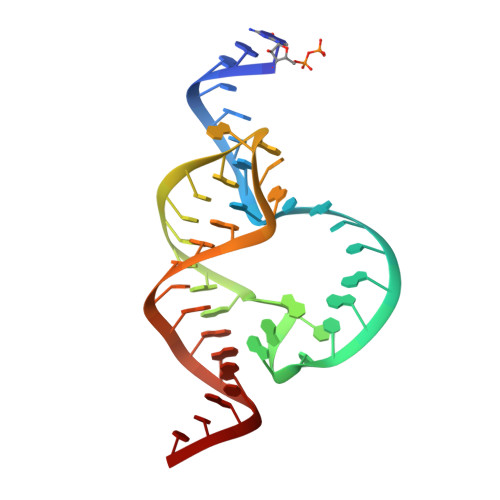
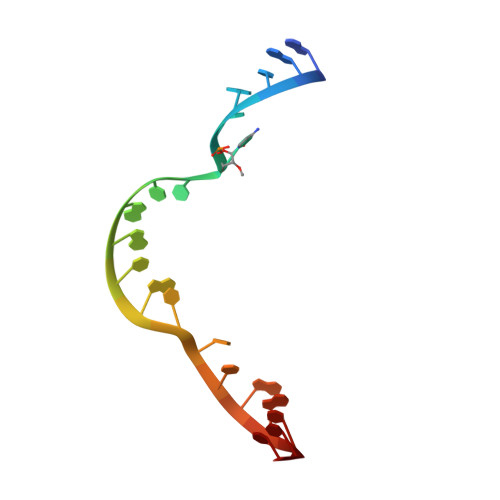
Active-Site Monovalent Cations Revealed in a 1.55 A Resolution Hammerhead Ribozyme Structure
Anderson, M., Schultz, E., Martick, M., Scott, W.G.(2013) J Mol Biology 425: 3790
- PubMed: 23711504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2013.05.017
- Primary Citation Related Structures:
3ZP8 - PubMed Abstract:
We have obtained a 1.55-Å crystal structure of a hammerhead ribozyme derived from Schistosoma mansoni under conditions that permit detailed observations of Na(+) ion binding in the ribozyme's active site. At least two such Na(+) ions are observed. The first Na(+) ion binds to the N7 of G10.1 and the adjacent A9 phosphate in a manner identical with that previously observed for divalent cations. A second Na(+) ion binds to the Hoogsteen face of G12, the general base in the hammerhead cleavage reaction, thereby potentially dissipating the negative charge of the catalytically active enolate form of the nucleotide base. A potential but more ambiguous third site bridges the A9 and scissile phosphates in a manner consistent with that of previous predictions. Hammerhead ribozymes have been observed to be active in the presence of high concentrations of monovalent cations, including Na(+), but the mechanism by which monovalent cations substitute for divalent cations in hammerhead catalysis remains unclear. Our results enable us to suggest that Na(+) directly and specifically substitutes for divalent cations in the hammerhead active site. The detailed geometry of the pre-catalytic active-site complex is also revealed with a new level of precision, thanks to the quality of the electron density maps obtained from what is currently the highest-resolution ribozyme structure in the Protein Data Bank.
- Department of Chemistry and Biochemistry and The Center for the Molecular Biology of RNA, 228 Sinsheimer Laboratories, University of California at Santa Cruz, Santa Cruz, CA 95064, USA.
Organizational Affiliation: