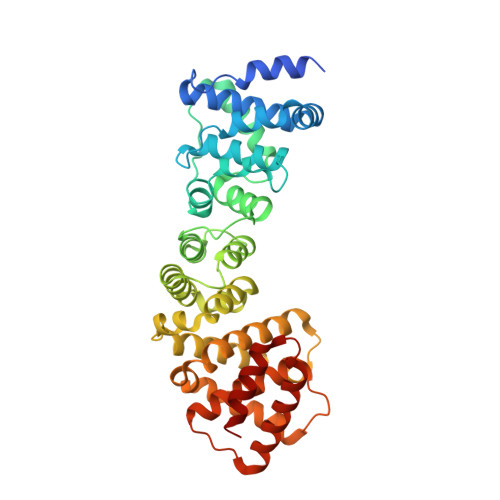
Structural Insights Into Regulation of Cohesion Establishment by Wpl1
Chatterjee, A., Zakian, S., Hu, X.-W., Singleton, M.R.(2013) EMBO J 32: 677
- PubMed: 23395900 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2013.16
- Primary Citation Related Structures:
3ZIK, 3ZIL - PubMed Abstract:
Correct segregation of duplicated chromosomes to daughter cells during mitosis requires the action of the cohesin complex. This tripartite ring-shaped molecule is involved in holding replicated sister chromatids together from S phase until anaphase onset. Establishment of stable cohesion involves acetylation of the Smc3 component of cohesin during replication by the Eco1 acetyltransferase. This has been proposed to antagonise the activity of another member of the cohesin complex, Wpl1. Here, we describe the X-ray structure of the conserved Wapl domain, and demonstrate that it binds the ATPase head of the Smc3 protein. We present data that suggest that Wpl1 may be involved in regulating the ATPase activity of cohesin, and that this may be subject to the acetylation state of Smc3. In addition, we present a structure of the Wapl domain bound to a functionally relevant segment of the Smc3 ATPase.
- Macromolecular Structure and Function Laboratory, Cancer Research UK, London Research Institute, London, UK.
Organizational Affiliation: