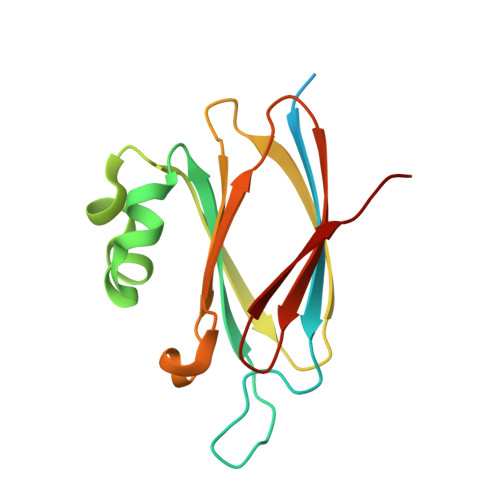
Structure of the T6Ss Lipoprotein Tssj1 from Pseudomonas Aeruginosa.
Robb, C.S., Assmus, M., Nano, F.E., Boraston, A.B.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 607
- PubMed: 23722835 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113012220
- Primary Citation Related Structures:
3ZHN - PubMed Abstract:
The type VI secretion system of Pseudomonas aeruginosa has been shown to be responsible for the translocation of bacteriolytic effectors into competing bacteria. A mechanistic understanding of this widely distributed secretion system is developing and structural studies of its components are ongoing. Two representative structures of one highly conserved component, TssJ, from Escherichia coli and Serratia marcescens have been published. Here, the X-ray crystal structure of TssJ1 from P. aeruginosa is presented at 1.4 Å resolution. The overall structure is conserved among the three proteins. This finding suggests that the homologues function in a similar manner and bolsters the understanding of the structure of this family of proteins.
- Department of Biochemistry and Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, British Columbia V8W 3P6, Canada.
Organizational Affiliation: