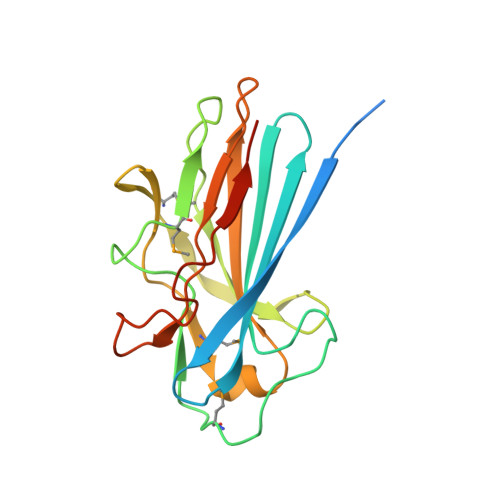
The Basic Keratin 10-Binding Domain of the Virulence-Associated Pneumococcal Serine-Rich Protein Psrp Adopts a Novel Mscramm Fold.
Schulte, T., Lofling, J., Mikaelsson, C., Kikhney, A., Hentrich, K., Diamante, A., Ebel, C., Normark, S., Svergun, D., Henriques-Normark, B., Achour, A.(2014) Open Biol 4: 0090
- PubMed: 24430336 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.130090
- Primary Citation Related Structures:
3ZGH, 3ZGI - PubMed Abstract:
Streptococcus pneumoniae is a major human pathogen, and a leading cause of disease and death worldwide. Pneumococcal invasive disease is triggered by initial asymptomatic colonization of the human upper respiratory tract. The pneumococcal serine-rich repeat protein (PsrP) is a lung-specific virulence factor whose functional binding region (BR) binds to keratin-10 (KRT10) and promotes pneumococcal biofilm formation through self-oligomerization. We present the crystal structure of the KRT10-binding domain of PsrP (BR187-385) determined to 2.0 Å resolution. BR187-385 adopts a novel variant of the DEv-IgG fold, typical for microbial surface components recognizing adhesive matrix molecules adhesins, despite very low sequence identity. An extended β-sheet on one side of the compressed, two-sided barrel presents a basic groove that possibly binds to the acidic helical rod domain of KRT10. Our study also demonstrates the importance of the other side of the barrel, formed by extensive well-ordered loops and stabilized by short β-strands, for interaction with KRT10.
- Science for Life Laboratory, Center for Infectious Medicine (CIM), Department of Medicine, Karolinska University Hospital Huddinge, Karolinska Institutet Science Park, Tomtebodavägen 23A Solna, Stockholm 17165, Sweden.
Organizational Affiliation: