Metal Sensing and Signal Transduction by Cnrx from Cupriavidus Metallidurans Ch34: Role of the Only Methionine Assessed by a Functional, Spectroscopic, and Theoretical Study
Trepreau, J., Grosse, C., Mouesca, J.-M., Sarret, G., Girard, E., Petit-Haertlein, I., Kuennemann, S., Desbourdes, C., De Rosny, E., Maillard, A.P., Nies, D.H., Coves, J.(2014) Metallomics 6: 263
- PubMed: 24154823
- DOI: https://doi.org/10.1039/c3mt00248a
- Primary Citation of Related Structures:
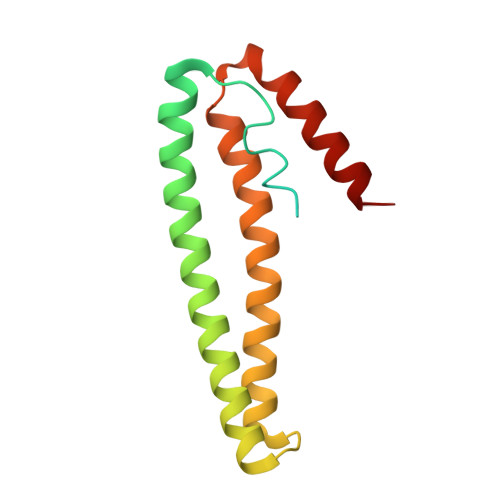
3ZG1 - PubMed Abstract:
When CnrX, the periplasmic sensor protein in the CnrYXH transmembrane signal transduction complex of Cupriavidus metallidurans CH34, binds the cognate metal ions Ni(II) or Co(II), the ECF-type sigma factor CnrH is made available in the cytoplasm for the RNA-polymerase to initiate transcription at the cnrYp and cnrCp promoters. Ni(II) or Co(II) are sensed by a metal-binding site with a N3O2S coordination sphere with octahedral geometry, where S stands for the thioether sulfur of the only methionine (Met123) residue of CnrX. The M123A-CnrX derivative has dramatically reduced signal propagation in response to metal sensing while the X-ray structure of Ni-bound M123A-CnrXs showed that the metal-binding site was not affected by the mutation. Ni(II) remained six-coordinate in M123A-CnrXs, with a water molecule replacing the sulfur as the sixth ligand. H32A-CnrXs, the soluble model of the wild-type membrane-anchored CnrX, was compared to the double mutants H32A-M123A-CnrXs and H32A-M123C-CnrXs to spectroscopically evaluate the role of this unique ligand in the binding site of Ni or Co. The Co- and Ni-bound forms of the protein display unusually blue-shifted visible spectra. TD-DFT calculations using structure-based models allowed identification and assignment of the electronic transitions of Co-bound form of the protein and its M123A derivative. Among them, the signature of the S-Co transition is distinguishable in the shoulder at 530 nm. In vitro affinity measurements point out the crucial role of Met123 in the selectivity for Ni or Co, and in vivo data support the conclusion that Met123 is a trigger of the signal transduction.
- Institut de Biologie Structurale, UMR 5075 CNRS-CEA-UJF-Grenoble-1, 6 Rue Jules Horowitz, 38042 Grenoble, France. jacques.coves@ibs.fr.
Organizational Affiliation: