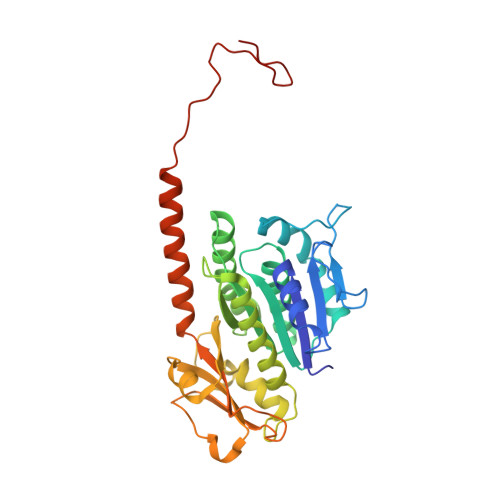
Structure of the Tubulin/Ftsz-Like Protein Tubz from Pseudomonas Bacteriophage Phikz
Aylett, C.H.S., Izore, T., Amos, L.A., Lowe, J.(2013) J Mol Biology 425: 2164
- PubMed: 23528827 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2013.03.019
- Primary Citation Related Structures:
3ZBP, 3ZBQ - PubMed Abstract:
Pseudomonas ΦKZ-like bacteriophages encode a group of related tubulin/FtsZ-like proteins believed to be essential for the correct centring of replicated bacteriophage virions within the bacterial host. In this study, we present crystal structures of the tubulin/FtsZ-like protein TubZ from Pseudomonas bacteriophage ΦKZ in both the monomeric and protofilament states, revealing that ΦKZ TubZ undergoes structural changes required to polymerise, forming a canonical tubulin/FtsZ-like protofilament. Combining our structures with previous work, we propose a polymerisation-depolymerisation cycle for the Pseudomonas bacteriophage subgroup of tubulin/FtsZ-like proteins. Electron cryo-microscopy of ΦKZ TubZ filaments polymerised in vitro implies a long-pitch helical arrangement for the constituent protofilaments. Intriguingly, this feature is shared by the other known subgroup of bacteriophage tubulin/FtsZ-like proteins from Clostridium species, which are thought to be involved in partitioning the genomes of bacteriophages adopting a pseudo-lysogenic life cycle.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: