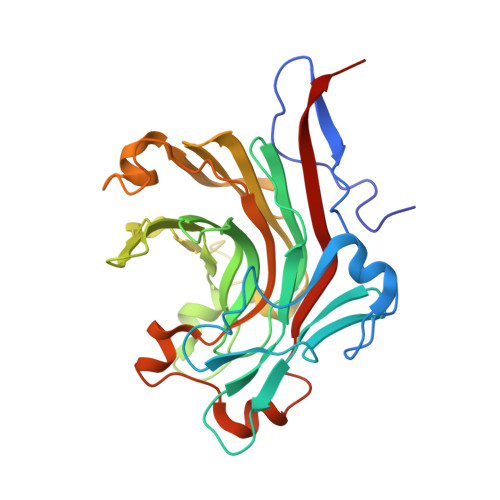
Crystal structure of the catalytic domain of a GH16 beta-agarase from a deep-sea bacterium, Microbulbifer thermotolerans JAMB-A94
Takagi, E., Hatada, Y., Akita, M., Ohta, Y., Yokoi, G., Miyazaki, T., Nishikawa, A., Tonozuka, T.(2015) Biosci Biotechnol Biochem 79: 625-632
- PubMed: 25483365
- DOI: https://doi.org/10.1080/09168451.2014.988680
- Primary Citation of Related Structures:
3WZ1 - PubMed Abstract:
A deep-sea bacterium, Microbulbifer thermotolerans JAMB-A94, has a β-agarase (MtAgaA) belonging to the glycoside hydrolase family (GH) 16. The optimal temperature of this bacterium for growth is 43-49 °C, and MtAgaA is stable at 60 °C, which is one of the most thermostable enzymes among GH16 β-agarases. Here, we determined the catalytic domain structure of MtAgaA. MtAgaA consists of a β-jelly roll fold, as observed in other GH16 enzymes. The structure of MtAgaA was most similar to two β-agarases from Zobellia galactanivorans, ZgAgaA, and ZgAgaB. Although the catalytic cleft structure of MtAgaA was similar to ZgAgaA and ZgAgaB, residues at subsite -4 of MtAgaA were not conserved between them. Also, an α-helix, designated as α4', was uniquely located near the catalytic cleft of MtAgaA. A comparison of the structures of the three enzymes suggested that multiple factors, including increased numbers of arginine and proline residues, could contribute to the thermostability of MtAgaA.
- a Japan Agency for Marine-Earth Science and Technology (JAMSTEC) , Yokosuka , Japan.
Organizational Affiliation: