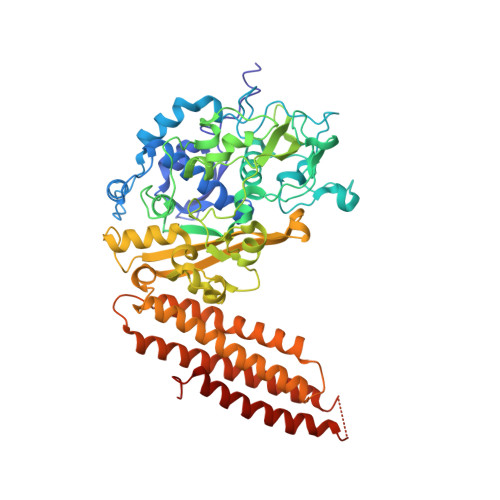
Crystal structure of Thermobifida fusca Cse1 reveals target DNA binding site.
Tay, M., Liu, S., Yuan, Y.A.(2015) Protein Sci 24: 236-245
- PubMed: 25420472 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2609
- Primary Citation Related Structures:
3WVO - PubMed Abstract:
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-CRISPR-associated (Cas) defense system is the only adaptive and inheritable immunity found in prokaryotes. The immunity is achieved through a multistep process of adaptation, expression, and interference. In the Type I-E system, interference is mediated by the CRISPR-associated complex for antiviral defense (Cascade), which recognizes invading double-stranded DNA (dsDNA) through the protospacer adjacent motif (PAM) by one of the Cascade components, Cse1. Here, we report the crystal structure of Thermobifida fusca Cse1 at 3.3 Å resolution. T. fusca Cse1 reveals the chair-like two-domain architecture with a well-defined flexible loop, L1, located at the larger N-terminal domain, which was not observed in previous structures of the single Cse1 protein. Structure-based mutagenesis analysis demonstrates that the well-defined flexible loop and a partially conserved structural motif ([FW]-X-[TH]) are involved in PAM binding and recognition, respectively. Moreover, structural docking of T. fusca Cse1 into Escherichia coli Cascade cryoelectron microscopy maps, coupled with structural comparison, reveals a conserved positive patch that is contiguous with Cse2 in the Cascade complex and adjacent to the Cas3 binding site, suggesting its role in R-loop formation/stabilization and the recruitment of Cas3 for target cleavage. Consistent with the structural observation, the introduction of alanine mutations at this positive patch abolished DNA binding activity by Cse1. Taken together, these results suggest that Cse1 is a critical Cascade component involved in Cascade assembly, dsDNA target recognition, R-loop formation, and Cas3 recruitment for target cleavage.
- Department of Biological Sciences and Centre for Bioimaging Sciences, National University of Singapore, Singapore, 117543, Singapore.
Organizational Affiliation: