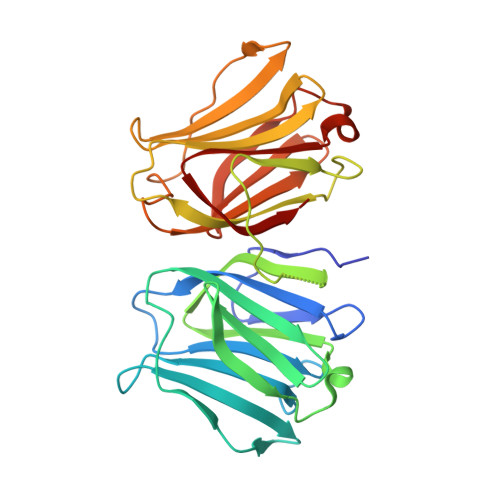
X-ray structure of a protease-resistant mutant form of human galectin-9 having two carbohydrate recognition domains with a metal-binding site
Yoshida, H., Nishi, N., Wada, K., Nakamura, T., Hirashima, M., Kuwabara, N., Kato, R., Kamitori, S.(2017) Biochem Biophys Res Commun 490: 1287-1293
- PubMed: 28687490 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.07.009
- Primary Citation Related Structures:
3WV6 - PubMed Abstract:
Galectin-9 (G9) is a tandem-repeat type β-galactoside-specific animal lectin having N-terminal and C-terminal carbohydrate recognition domains (N-CRD and C-CRD, respectively) joined by a linker peptide that is involved in the immune system. G9 is divalent in glycan binding, and structural information about the spatial arrangement of the two CRDs is very important for elucidating its biological functions. As G9 is protease sensitive due to the long linker, the protease-resistant mutant form of G9 (G9Null) was developed by modification of the linker peptide, while retaining its biological functions. The X-ray structure of a mutant form of G9Null with the replacement of Arg221 by Ser (G9Null_R221S) having two CRDs was determined. The structure of G9Null_R221S was compact to associate the two CRDs in the back-to-back orientation with a large interface area, including hydrogen bonds and hydrophobic interactions. A metal ion was newly found in the galectin structure, possibly contributing to the stable structure of protein. The presented X-ray structure was thought to be one of the stable structures of G9, which likely occurs in solution. This was supported by structural comparisons with other tandem-repeated galectins and the analyses of protein thermostability by CD spectra measurements.
- Life Science Research Center and Faculty of Medicine, Kagawa University, 1750-1, Ikenobe, Miki-cho, Kita-gun, Kagawa 761-0793, Japan.
Organizational Affiliation: