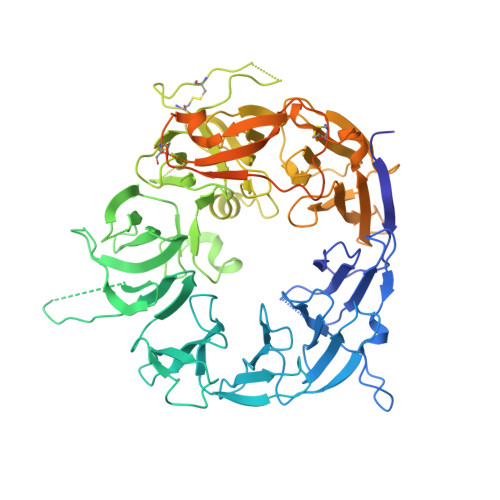
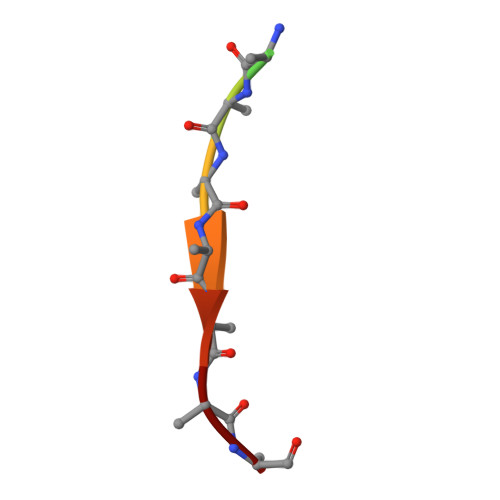
Structural basis for amyloidogenic peptide recognition by sorLA.
Kitago, Y., Nagae, M., Nakata, Z., Yagi-Utsumi, M., Takagi-Niidome, S., Mihara, E., Nogi, T., Kato, K., Takagi, J.(2015) Nat Struct Mol Biol 22: 199-206
- PubMed: 25643321 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2954
- Primary Citation Related Structures:
3WSX, 3WSY, 3WSZ - PubMed Abstract:
SorLA is a neuronal sorting receptor considered to be a major risk factor for Alzheimer's disease. We have recently reported that it directs lysosomal targeting of nascent neurotoxic amyloid-β (Aβ) peptides by directly binding Aβ. Here, we determined the crystal structure of the human sorLA domain responsible for Aβ capture, Vps10p, in an unbound state and in complex with two ligands. Vps10p assumes a ten-bladed β-propeller fold with a large tunnel at the center. An internal ligand derived from the sorLA propeptide bound inside the tunnel to extend the β-sheet of one of the propeller blades. The structure of the sorLA Vps10p-Aβ complex revealed that the same site is used. Peptides are recognized by sorLA Vps10p in redundant modes without strict dependence on a particular amino acid sequence, thus suggesting a broad specificity toward peptides with a propensity for β-sheet formation.
- Institute for Protein Research, Osaka University, Suita, Japan.
Organizational Affiliation: