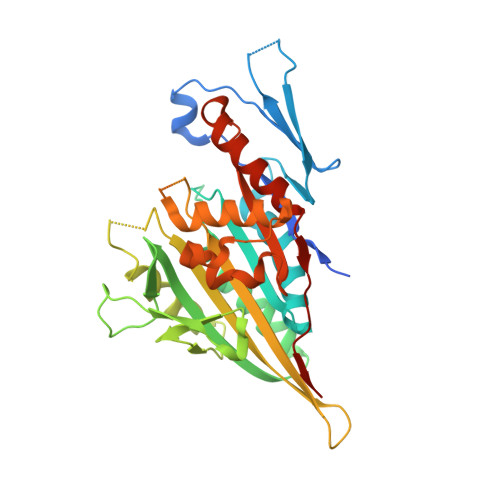
Structural basis of new allosteric inhibition in Kinesin spindle protein eg5
Yokoyama, H., Sawada, J., Katoh, S., Matsuno, K., Ogo, N., Ishikawa, Y., Hashimoto, H., Fujii, S., Asai, A.(2015) ACS Chem Biol 10: 1128-1136
- PubMed: 25622007 Search on PubMed
- DOI: https://doi.org/10.1021/cb500939x
- Primary Citation Related Structures:
3WPN - PubMed Abstract:
Kinesin spindle protein Eg5 is a target for anticancer therapies, and small molecule inhibitors of its ATPase activity have been developed. We herein report for the first time the crystal structure of and biochemical studies on the Eg5 motor domain in complex with a new type of allosteric inhibitor. The biphenyl-type inhibitor PVZB1194 binds to the α4/α6 allosteric pocket 15 Å from the ATP-binding pocket, which differs from conventional allosteric inhibitors that bind to the allosteric L5/α2/α3 pocket of Eg5. Binding of the inhibitor is involved in the neck-linker conformation and also causes conformational changes around the ATP-binding pocket through Tyr104 to affect the interaction of ATP with the pocket. This structure provides useful information for the development of novel types of allosteric drugs as well as a novel insight into the molecular mechanism responsible for regulating the motor activity of kinesins.
- †Department of Physical Biochemistry, School of Pharmaceutical Sciences, University of Shizuoka, Shizuoka 422-8526, Japan.
Organizational Affiliation: