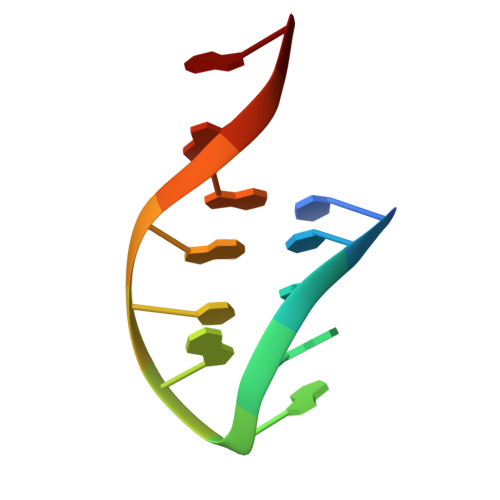
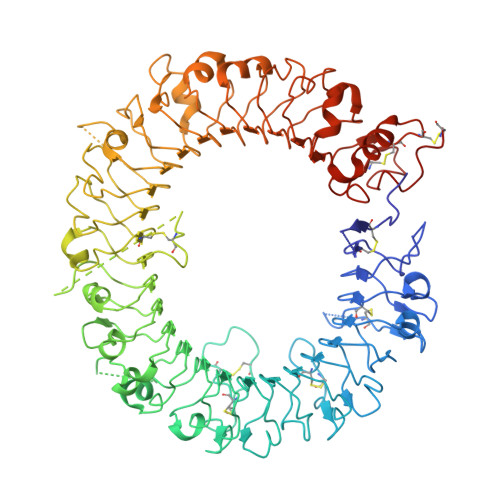
Structural basis of CpG and inhibitory DNA recognition by Toll-like receptor 9
Ohto, U., Shibata, T., Tanji, H., Ishida, H., Krayukhina, E., Uchiyama, S., Miyake, K., Shimizu, T.(2015) Nature 520: 702-705
- PubMed: 25686612
- DOI: https://doi.org/10.1038/nature14138
- Primary Citation of Related Structures:
3WPB, 3WPC, 3WPD, 3WPE, 3WPF, 3WPG, 3WPH, 3WPI - PubMed Abstract:
Innate immunity serves as the first line of defence against invading pathogens such as bacteria and viruses. Toll-like receptors (TLRs) are examples of innate immune receptors, which sense specific molecular patterns from pathogens and activate immune responses. TLR9 recognizes bacterial and viral DNA containing the cytosine-phosphate-guanine (CpG) dideoxynucleotide motif. The molecular basis by which CpG-containing DNA (CpG-DNA) elicits immunostimulatory activity via TLR9 remains to be elucidated. Here we show the crystal structures of three forms of TLR9: unliganded, bound to agonistic CpG-DNA, and bound to inhibitory DNA (iDNA). Agonistic-CpG-DNA-bound TLR9 formed a symmetric TLR9-CpG-DNA complex with 2:2 stoichiometry, whereas iDNA-bound TLR9 was a monomer. CpG-DNA was recognized by both protomers in the dimer, in particular by the amino-terminal fragment (LRRNT-LRR10) from one protomer and the carboxy-terminal fragment (LRR20-LRR22) from the other. The iDNA, which formed a stem-loop structure suitable for binding by intramolecular base pairing, bound to the concave surface from LRR2-LRR10. This structure serves as an important basis for improving our understanding of the functional mechanisms of TLR9.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: