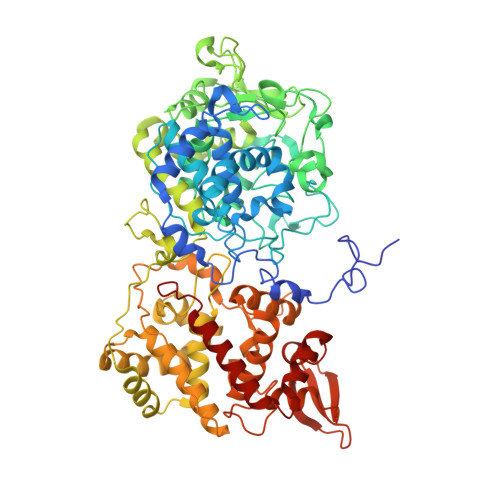
The 2.2 angstrom resolution structure of the catalase-peroxidase KatG from Synechococcus elongatus PCC7942.
Kamachi, S., Wada, K., Tamoi, M., Shigeoka, S., Tada, T.(2014) Acta Crystallogr F Struct Biol Commun 70: 288-293
- PubMed: 24598912 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14002052
- Primary Citation Related Structures:
3WNU - PubMed Abstract:
The crystal structure of catalase-peroxidase from Synechococcus elongatus PCC7942 (SeKatG) was solved by molecular replacement and refined to an Rwork of 16.8% and an Rfree of 20.6% at 2.2 Å resolution. The asymmetric unit consisted of only one subunit of the catalase-peroxidase molecule, including a protoporphyrin IX haem moiety and two sodium ions. A typical KatG covalent adduct was formed, Met248-Tyr222-Trp94, which is a key structural element for catalase activity. The crystallographic equivalent subunit was created by a twofold symmetry operation to form the functional dimer. The overall structure of the dimer was quite similar to other KatGs. One sodium ion was located close to the proximal Trp314. The location and configuration of the proximal cation site were very similar to those of typical peroxidases such as ascorbate peroxidase. These features may provide a structural basis for the behaviour of the radical localization/delocalization during the course of the enzymatic reaction.
- School of Graduate Science, Osaka Prefecture University, Sakai, Osaka 599-8531, Japan.
Organizational Affiliation: