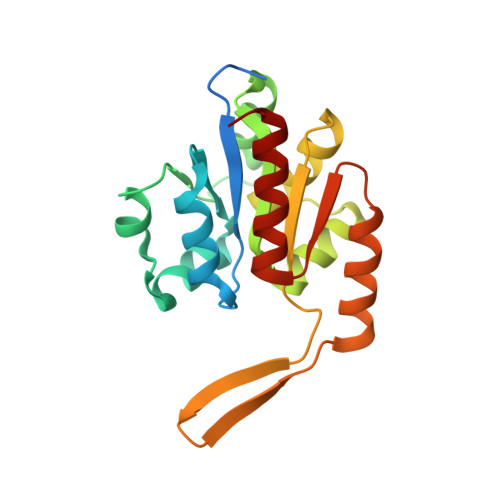
Structure of dihydromethanopterin reductase, a cubic protein cage for redox transfer
Mcnamara, D.E., Cascio, D., Jorda, J., Bustos, C., Wang, T.C., Rasche, M.E., Yeates, T.O., Bobik, T.A.(2014) J Biological Chem 289: 8852-8864
- PubMed: 24523405
- DOI: https://doi.org/10.1074/jbc.M113.522342
- Primary Citation Related Structures:
3WIS, 4MWG - PubMed Abstract:
Dihydromethanopterin reductase (Dmr) is a redox enzyme that plays a key role in generating tetrahydromethanopterin (H4MPT) for use in one-carbon metabolism by archaea and some bacteria. DmrB is a bacterial enzyme understood to reduce dihydromethanopterin (H2MPT) to H4MPT using flavins as the source of reducing equivalents, but the mechanistic details have not been elucidated previously. Here we report the crystal structure of DmrB from Burkholderia xenovorans at a resolution of 1.9 Å. Unexpectedly, the biological unit is a 24-mer composed of eight homotrimers located at the corners of a cubic cage-like structure. Within a homotrimer, each monomer-monomer interface exhibits an active site with two adjacently bound flavin mononucleotide (FMN) ligands, one deeply buried and tightly bound and one more peripheral, for a total of 48 ligands in the biological unit. Computational docking suggested that the peripheral site could bind either the observed FMN (the electron donor for the overall reaction) or the pterin, H2MPT (the electron acceptor for the overall reaction), in configurations ideal for electron transfer to and from the tightly bound FMN. On this basis, we propose that DmrB uses a ping-pong mechanism to transfer reducing equivalents from FMN to the pterin substrate. Sequence comparisons suggested that the catalytic mechanism is conserved among the bacterial homologs of DmrB and partially conserved in archaeal homologs, where an alternate electron donor is likely used. In addition to the mechanistic revelations, the structure of DmrB could help guide the development of anti-obesity drugs based on modification of the ecology of the human gut.
- From the Department of Chemistry and Biochemistry.
Organizational Affiliation: