Crystal Structure of Clostridium botulinum Whole Hemagglutinin Reveals a Huge Triskelion-shaped Molecular Complex
Amatsu, S., Sugawara, Y., Matsumura, T., Kitadokoro, K., Fujinaga, Y.(2013) J Biological Chem 288: 35617-35625
- PubMed: 24165130 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.521179
- Primary Citation Related Structures:
3WIN - PubMed Abstract:
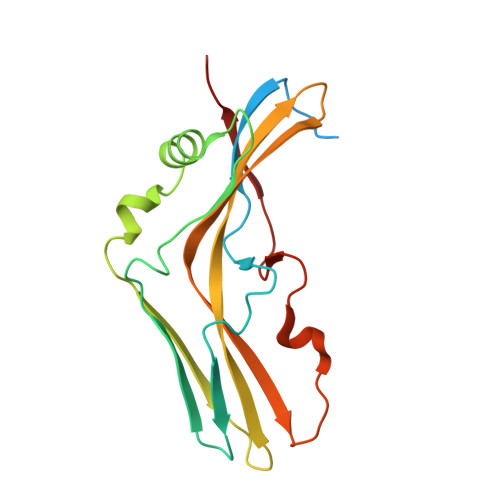
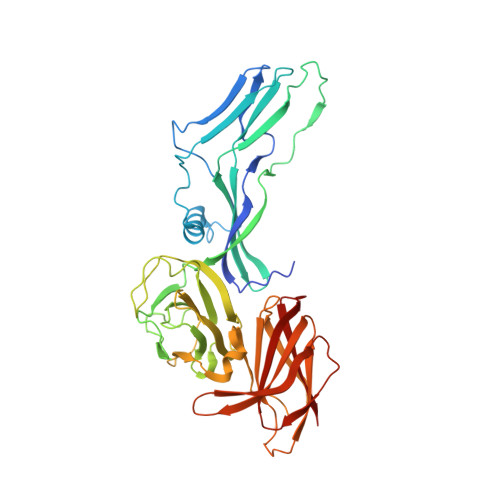
Clostridium botulinum HA is a component of the large botulinum neurotoxin complex and is critical for its oral toxicity. HA plays multiple roles in toxin penetration in the gastrointestinal tract, including protection from the digestive environment, binding to the intestinal mucosal surface, and disruption of the epithelial barrier. At least two properties of HA contribute to these roles: the sugar-binding activity and the barrier-disrupting activity that depends on E-cadherin binding of HA. HA consists of three different proteins, HA1, HA2, and HA3, whose structures have been partially solved and are made up mainly of β-strands. Here, we demonstrate structural and functional reconstitution of whole HA and present the complete structure of HA of serotype B determined by x-ray crystallography at 3.5 Å resolution. This structure reveals whole HA to be a huge triskelion-shaped molecule. Our results suggest that whole HA is functionally and structurally separable into two parts: HA1, involved in recognition of cell-surface carbohydrates, and HA2-HA3, involved in paracellular barrier disruption by E-cadherin binding.
- From the Graduate School of Science and Technology, Department of Biomolecular Engineering, Kyoto Institute of Technology, Matsugasakigosyokaido-cho, Sakyo-ku, Kyoto 606-8585 and.
Organizational Affiliation: