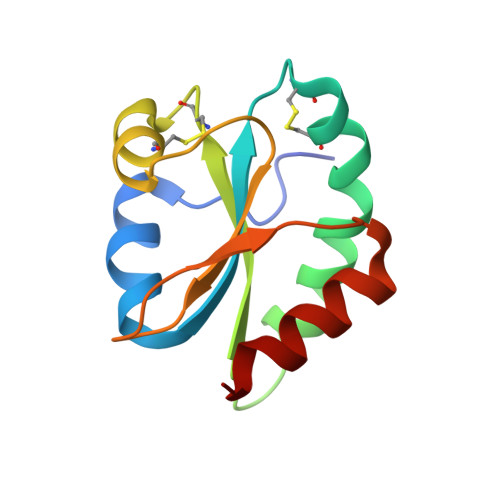
Radically different thioredoxin domain arrangement of ERp46, an efficient disulfide bond introducer of the mammalian PDI family
Kojima, R., Okumura, M., Masui, S., Kanemura, S., Inoue, M., Saiki, M., Yamaguchi, H., Hikima, T., Suzuki, M., Akiyama, S., Inaba, K.(2014) Structure 22: 431-443
- PubMed: 24462249 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2013.12.013
- Primary Citation Related Structures:
3WGD, 3WGE, 3WGX - PubMed Abstract:
The mammalian endoplasmic reticulum (ER) contains a diverse oxidative protein folding network in which ERp46, a member of the protein disulfide isomerase (PDI) family, serves as an efficient disulfide bond introducer together with Peroxiredoxin-4 (Prx4). We revealed a radically different molecular architecture of ERp46, in which the N-terminal two thioredoxin (Trx) domains with positively charged patches near their peptide-binding site and the C-terminal Trx are linked by unusually long loops and arranged extendedly, forming an opened V-shape. Whereas PDI catalyzes native disulfide bond formation by the cooperative action of two mutually facing redox-active sites on folding intermediates bound to the central cleft, ERp46 Trx domains are separated, act independently, and engage in rapid but promiscuous disulfide bond formation during early oxidative protein folding. Thus, multiple PDI family members likely contribute to different stages of oxidative folding and work cooperatively to ensure the efficient production of multi-disulfide proteins in the ER.
- Division of Protein Chemistry, Medical Institute of Bioregulation, Kyushu University, 3-1-1 Maidashi, Higashi-ku, Fukuoka 812-8582, Japan.
Organizational Affiliation: