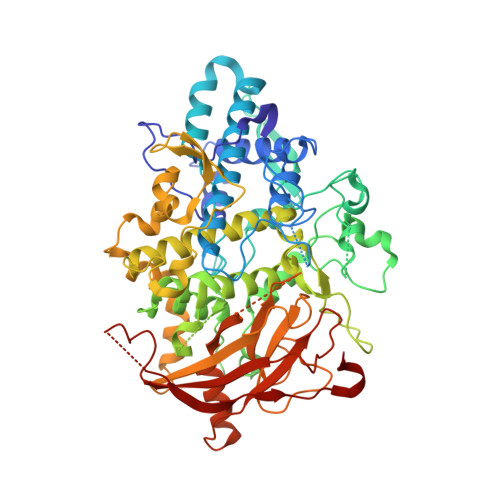
Crystal structures of copper-depleted and copper-bound fungal pro-tyrosinase: insights into endogenous cysteine-dependent copper incorporation.
Fujieda, N., Yabuta, S., Ikeda, T., Oyama, T., Muraki, N., Kurisu, G., Itoh, S.(2013) J Biological Chem 288: 22128-22140
- PubMed: 23749993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.477612
- Primary Citation Related Structures:
3W6Q, 3W6W - PubMed Abstract:
Tyrosinase, a dinuclear copper monooxygenase/oxidase, plays a crucial role in the melanin pigment biosynthesis. The structure and functions of tyrosinase have so far been studied extensively, but the post-translational maturation process from the pro-form to the active form has been less explored. In this study, we provide the crystal structures of Aspergillus oryzae full-length pro-tyrosinase in the holo- and the apo-forms at 1.39 and 2.05 Å resolution, respectively, revealing that Phe(513) on the C-terminal domain is accommodated in the substrate-binding site as a substrate analog to protect the dicopper active site from substrate access (proteolytic cleavage of the C-terminal domain or deformation of the C-terminal domain by acid treatment transforms the pro-tyrosinase to the active enzyme (Fujieda, N., Murata, M., Yabuta, S., Ikeda, T., Shimokawa, C., Nakamura, Y., Hata, Y., and Itoh, S. (2012) ChemBioChem. 13, 193-201 and Fujieda, N., Murata, M., Yabuta, S., Ikeda, T., Shimokawa, C., Nakamura, Y., Hata, Yl, and Itoh, S. (2013) J. Biol. Inorg. Chem. 18, 19-26). Detailed crystallographic analysis and structure-based mutational studies have shown that the copper incorporation into the active site is governed by three cysteines as follows: Cys(92), which is covalently bound to His(94) via an unusual thioether linkage in the holo-form, and Cys(522) and Cys(525) of the CXXC motif located on the C-terminal domain. Molecular mechanisms of the maturation processes of fungal tyrosinase involving the accommodation of the dinuclear copper unit, the post-translational His-Cys thioether cross-linkage formation, and the proteolytic C-terminal cleavage to produce the active tyrosinase have been discussed on the basis of the detailed structural information.
- Department of Material and Life Science, Division of Advanced Science and Biotechnology, Graduate School of Engineering, Osaka University, 2-1 Yamada-oka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: