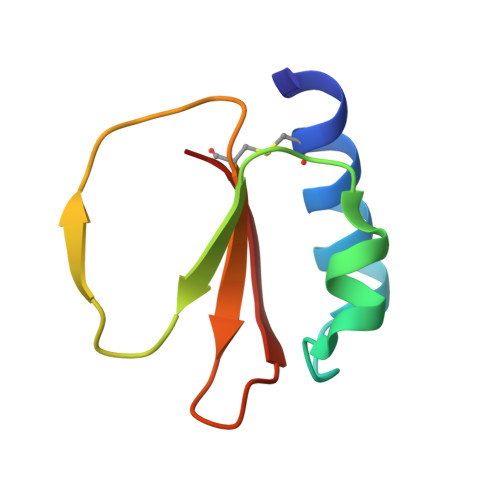
X-ray Structure Analysis and Characterization of AFUEI, an Elastase Inhibitor from Aspergillus fumigatus
Sakuma, M., Imada, K., Okumura, Y., Uchiya, K., Yamashita, N., Ogawa, K., Hijikata, A., Shirai, T., Homma, M., Nikai, T.(2013) J Biological Chem 288: 17451-17459
- PubMed: 23640894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.433987
- Primary Citation Related Structures:
3W0D, 3W0E - PubMed Abstract:
Elastase from Aspergillus sp. is an important factor for aspergillosis. AFUEI is an inhibitor of the elastase derived from Aspergillus fumigatus. AFUEI is a member of the I78 inhibitor family and has a high inhibitory activity against elastases of Aspergillus fumigatus and Aspergillus flavus, human neutrophil elastase and bovine chymotrypsin, but does not inhibit bovine trypsin. Here we report the crystal structure of AFUEI in two crystal forms. AFUEI is a wedge-shaped protein composed of an extended loop and a scaffold protein core. The structure of AFUEI shows remarkable similarity to serine protease inhibitors of the potato inhibitor I family, although they are classified into different inhibitor families. A structural comparison with the potato I family inhibitors suggests that the extended loop of AFUEI corresponds to the binding loop of the potato inhibitor I family, and AFUEI inhibits its cognate proteases through the same mechanism as the potato I family inhibitors.
- Division of Biological Science, Graduate School of Science, Nagoya University, Furo-cho, Chikusa-Ku, Nagoya 464-8602, Japan.
Organizational Affiliation: