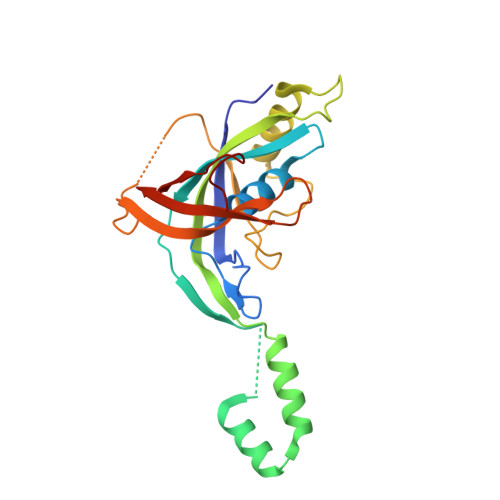
Conservation and Variability in the Structure and Function of the Cas5d Endoribonuclease in the CRISPR-Mediated Microbial Immune System
Koo, Y., Ka, D., Kim, E.J., Suh, N., Bae, E.(2013) J Mol Biology 425: 3799-3810
- PubMed: 23500492
- DOI: https://doi.org/10.1016/j.jmb.2013.02.032
- Primary Citation Related Structures:
3VZH, 3VZI - PubMed Abstract:
Clustered regularly interspaced short palindromic repeats (CRISPRs) and CRISPR-associated (Cas) proteins form an RNA-mediated microbial immune system against invading foreign genetic elements. Cas5 proteins constitute one of the most prevalent Cas protein families in CRISPR-Cas systems and are predicted to have RNA recognition motif (RRM) domains. Cas5d is a subtype I-C-specific Cas5 protein that can be divided into two distinct subgroups, one of which has extra C-terminal residues while the other contains a longer insertion in the middle of its N-terminal RRM domain. Here, we report crystal structures of Cas5d from Streptococcus pyogenes and Xanthomonas oryzae, which respectively represent the two Cas5d subgroups. Despite a common domain architecture consisting of an N-terminal RRM domain and a C-terminal β-sheet domain, the structural differences between the two Cas5d proteins are highlighted by the presence of a unique extended helical region protruding from the N-terminal RRM domain of X. oryzae Cas5d. We also demonstrate that Cas5d proteins possess not only specific endoribonuclease activity for CRISPR RNAs but also nonspecific double-stranded DNA binding affinity. These findings suggest that Cas5d may play multiple roles in CRISPR-mediated immunity. Furthermore, the specific RNA processing was also observed between S. pyogenes Cas5d protein and X. oryzae CRISPR RNA and vice versa. This cross-species activity of Cas5d provides a special opportunity for elucidating conserved features of the CRISPR RNA processing event.
- Department of Agricultural Biotechnology, Seoul National University, Seoul 151-921, Korea.
Organizational Affiliation: