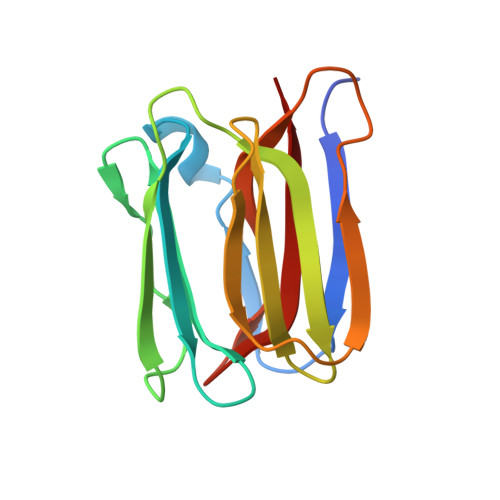
Structural Basis for Multiple Sugar Recognition of Jacalin-related Human ZG16p Lectin
Kanagawa, M., Liu, Y., Hanashima, S., Ikeda, A., Chai, W., Nakano, Y., Kojima-Aikawa, K., Feizi, T., Yamaguchi, Y.(2014) J Biological Chem 289: 16954-16965
- PubMed: 24790092
- DOI: https://doi.org/10.1074/jbc.M113.539114
- Primary Citation of Related Structures:
3VY6, 3VY7, 3VZE, 3VZF, 3VZG - PubMed Abstract:
ZG16p is a soluble mammalian lectin, the first to be described with a Jacalin-related β-prism-fold. ZG16p has been reported to bind both to glycosaminoglycans and mannose. To determine the structural basis of the multiple sugar-binding properties, we conducted glycan microarray analyses of human ZG16p. We observed that ZG16p preferentially binds to α-mannose-terminating short glycans such as Ser/Thr-linked O-mannose, but not to high mannose-type N-glycans. Among sulfated glycosaminoglycan oligomers examined, chondroitin sulfate B and heparin oligosaccharides showed significant binding. Crystallographic studies of human ZG16p lectin in the presence of selected ligands revealed the mechanism of multiple sugar recognition. Manα1-3Man and Glcβ1-3Glc bound in different orientations: the nonreducing end of the former and the reducing end of the latter fitted in the canonical shallow mannose binding pocket. Solution NMR analysis using (15)N-labeled ZG16p defined the heparin-binding region, which is on an adjacent flat surface of the protein. On-array competitive binding assays suggest that it is possible for ZG16p to bind simultaneously to both types of ligands. Recognition of a broad spectrum of ligands by ZG16p may account for the multiple functions of this lectin in the formation of zymogen granules via glycosaminoglycan binding, and in the recognition of pathogens in the digestive system through α-mannose-related recognition.
- From the Structural Glycobiology Team, Systems Glycobiology Research Group, RIKEN-Max Planck Joint Research Center, RIKEN Global Research Cluster, 2-1 Hirosawa, Wako, Saitama 351-0198, Japan.
Organizational Affiliation: