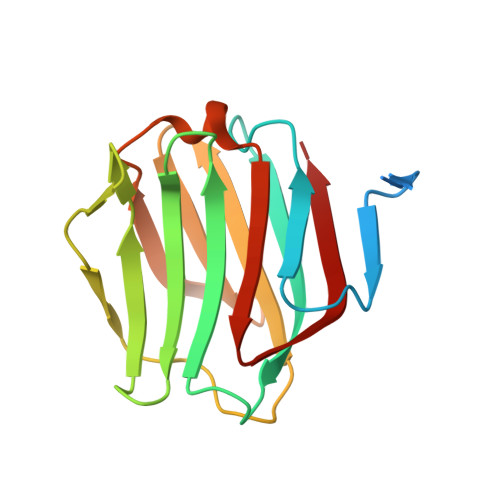
Structural basis of preferential binding of fucose-containing saccharide by the Caenorhabditis elegans galectin LEC-6
Makyio, H., Takeuchi, T., Tamura, M., Nishiyama, K., Takahashi, H., Natsugari, H., Arata, Y., Kasai, K., Yamada, Y., Wakatsuki, S., Kato, R.(2013) Glycobiology 23: 797-805
- PubMed: 23481096 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwt017
- Primary Citation Related Structures:
3VV1 - PubMed Abstract:
Galectins are a group of lectins that can bind carbohydrate chains containing β-galactoside units. LEC-6, a member of galectins of Caenorhabditis elegans, binds fucose-containing saccharides. We solved the crystal structure of LEC-6 in complex with galactose-β1,4-fucose (Galβ1-4Fuc) at 1.5 Å resolution. The overall structure of the protein and the identities of the amino-acid residues binding to the disaccharide are similar to those of other galectins. However, further structural analysis and multiple sequence alignment between LEC-6 and other galectins indicate that a glutamic acid residue (Glu67) is important for the preferential binding between LEC-6 and the fucose moiety of the Galβ1-4Fuc unit. Frontal affinity chromatography analysis indicated that the affinities of E67D and E67A mutants for Galβ1-4Fuc are lower than that of wild-type LEC-6. Furthermore, the affinities of Glu67 mutants for an endogenous oligosaccharide, which contains a Galβ1-4Fuc unit, are drastically reduced relative to that of the wild-type protein. We conclude that the Glu67 in the oligosaccharide-binding site assists the recognition of the fucose moiety by LEC-6.
- Structural Biology Research Center, Photon Factory, Institute of Materials Structure Science, High Energy Accelerator Research Organization KEK, 1-1 Oho, Tsukuba, Ibaraki 305-0801, Japan.
Organizational Affiliation: