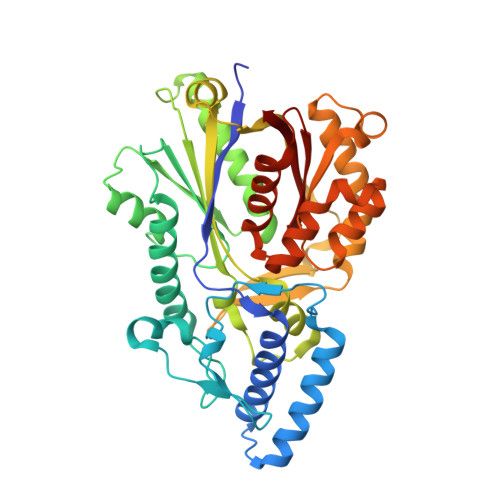
Structural basis for cyclization specificity of two Azotobacter type III polyketide synthases: a single amino acid substitution reverses their cyclization specificity
Satou, R., Miyanaga, A., Ozawa, H., Funa, N., Katsuyama, Y., Miyazono, K., Tanokura, M., Ohnishi, Y., Horinouchi, S.(2013) J Biological Chem 288: 34146-34157
- PubMed: 24100027
- DOI: https://doi.org/10.1074/jbc.M113.487272
- Primary Citation Related Structures:
3VS8, 3VS9 - PubMed Abstract:
Type III polyketide synthases (PKSs) show diverse cyclization specificity. We previously characterized two Azotobacter type III PKSs (ArsB and ArsC) with different cyclization specificity. ArsB and ArsC, which share a high sequence identity (71%), produce alkylresorcinols and alkylpyrones through aldol condensation and lactonization of the same polyketomethylene intermediate, respectively. Here we identified a key amino acid residue for the cyclization specificity of each enzyme by site-directed mutagenesis. Trp-281 of ArsB corresponded to Gly-284 of ArsC in the amino acid sequence alignment. The ArsB W281G mutant synthesized alkylpyrone but not alkylresorcinol. In contrast, the ArsC G284W mutant synthesized alkylresorcinol with a small amount of alkylpyrone. These results indicate that this amino acid residue (Trp-281 of ArsB or Gly-284 of ArsC) should occupy a critical position for the cyclization specificity of each enzyme. We then determined crystal structures of the wild-type and G284W ArsC proteins at resolutions of 1.76 and 1.99 Å, respectively. Comparison of these two ArsC structures indicates that the G284W substitution brings a steric wall to the active site cavity, resulting in a significant reduction of the cavity volume. We postulate that the polyketomethylene intermediate can be folded to a suitable form for aldol condensation only in such a relatively narrow cavity of ArsC G284W (and presumably ArsB). This is the first report on the alteration of cyclization specificity from lactonization to aldol condensation for a type III PKS. The ArsC G284W structure is significant as it is the first reported structure of a microbial resorcinol synthase.
- Department of Biotechnology, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: