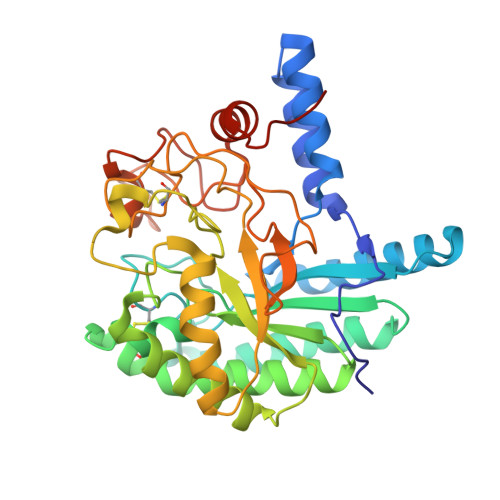
Comparison of the structural changes in two cellobiohydrolases, CcCel6A and CcCel6C, from Coprinopsis cinerea - a tweezer-like motion in the structure of CcCel6C
Tamura, M., Miyazaki, T., Tanaka, Y., Yoshida, M., Nishikawa, A., Tonozuka, T.(2012) FEBS J 279: 1871-1882
- PubMed: 22429290 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08568.x
- Primary Citation Related Structures:
3VOF, 3VOG, 3VOH, 3VOI, 3VOJ - PubMed Abstract:
The basidiomycete Coprinopsis cinerea produces five cellobiohydrolases belonging to glycoside hydrolase family 6 (GH6). Among these enzymes, C. cinerea cellulase 6C (CcCel6C), but not C. cinerea cellulase 6A (CcCel6A), can efficiently hydrolyze carboxymethyl cellulose and is constitutively expressed in C. cinerea. In contrast, CcCel6A possesses a cellulose-binding domain, and is strongly induced by cellobiose. Here, we determined the crystal structures of the CcCel6A catalytic domain complexed with a Hepes buffer molecule, with cellobiose, and with p-nitrophenyl β-D-cellotrioside (pNPG3). A notable feature of the GH6 cellobiohydrolases is that the active site is enclosed by two loops to form a tunnel, and the loops have been demonstrated to open and close in response to ligand binding. The enclosed tunnel of CcCel6A-Hepes is seen as the open form, whereas the tunnels of CcCel6A-cellobiose and CcCel6A-pNPG3 adopt the closed form. pNPG3 was not hydrolyzed by CcCel6A, and bound in subsites +1 to +4. On the basis of this observation, we constructed two mutants, CcCel6A D164A and CcCel6C D102A. Neither CcCel6A D164A nor CcCel6C D102A hydrolyze phosphoric acid-swollen cellulose. We have previously determined the crystal structures of CcCel6C unbound and in complex with ligand, both of which adopt the open form. In the present study, both CcCel6A and CcCel6C mutants were identified as the closed form. However, the motion angle of CcCel6C was more than 10-fold greater than that of CcCel6A. The width of the active site cleft of CcCel6C was narrowed, owing to a tweezer-like motion.
- Department of Applied Biological Science, Tokyo University of Agriculture and Technology, Tokyo, Japan.
Organizational Affiliation: