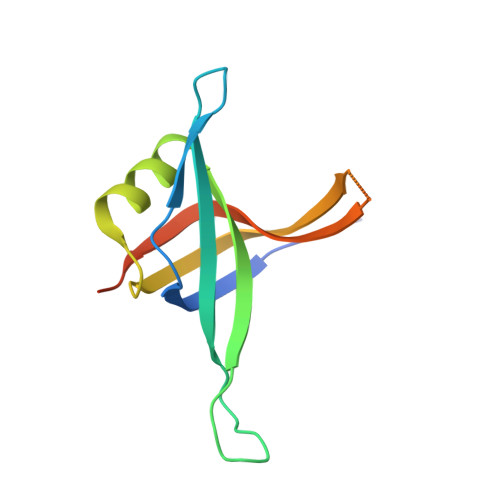
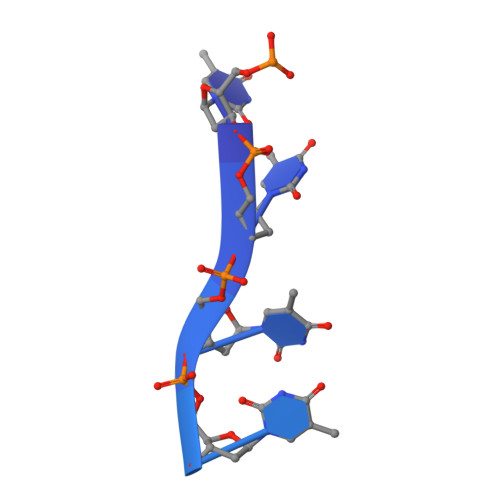
Genetic recombination in Bacillus subtilis: a division of labor between two single-strand DNA-binding proteins.
Yadav, T., Carrasco, B., Myers, A.R., George, N.P., Keck, J.L., Alonso, J.C.(2012) Nucleic Acids Res 40: 5546-5559
- PubMed: 22373918
- DOI: https://doi.org/10.1093/nar/gks173
- Primary Citation of Related Structures:
3VDY - PubMed Abstract:
We have investigated the structural, biochemical and cellular roles of the two single-stranded (ss) DNA-binding proteins from Bacillus subtilis, SsbA and SsbB. During transformation, SsbB localizes at the DNA entry pole where it binds and protects internalized ssDNA. The 2.8-Å resolution structure of SsbB bound to ssDNA reveals a similar overall protein architecture and ssDNA-binding surface to that of Escherichia coli SSB. SsbA, which binds ssDNA with higher affinity than SsbB, co-assembles onto SsbB-coated ssDNA and the two proteins inhibit ssDNA binding by the recombinase RecA. During chromosomal transformation, the RecA mediators RecO and DprA provide RecA access to ssDNA. Interestingly, RecO interaction with ssDNA-bound SsbA helps to dislodge both SsbA and SsbB from the DNA more efficiently than if the DNA is coated only with SsbA. Once RecA is nucleated onto the ssDNA, RecA filament elongation displaces SsbA and SsbB and enables RecA-mediated DNA strand exchange. During plasmid transformation, RecO localizes to the entry pole and catalyzes annealing of SsbA- or SsbA/SsbB-coated complementary ssDNAs to form duplex DNA with ssDNA tails. Our results provide a mechanistic framework for rationalizing the coordinated events modulated by SsbA, SsbB and RecO that are crucial for RecA-dependent chromosomal transformation and RecA-independent plasmid transformation.
- Departamento de Biotecnología Microbiana, Centro Nacional de Biotecnología, CSIC, 28049 Madrid, Spain.
Organizational Affiliation: