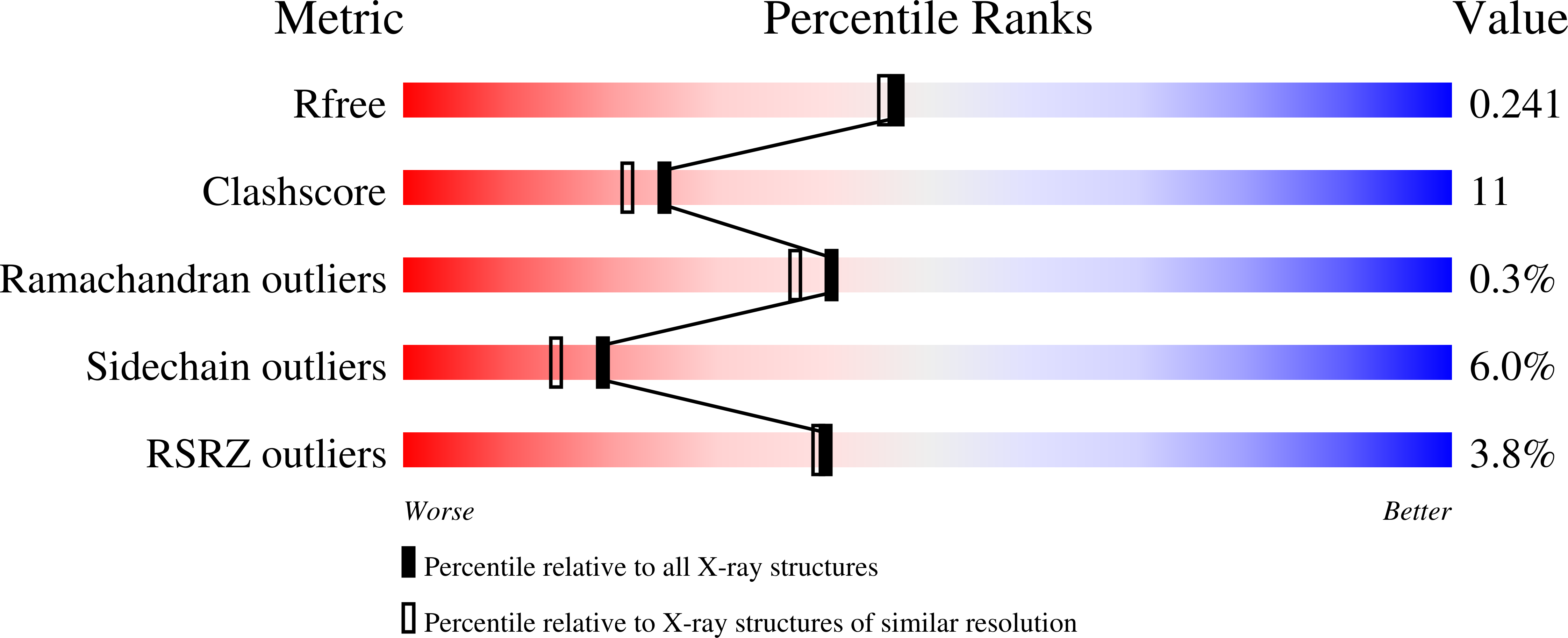
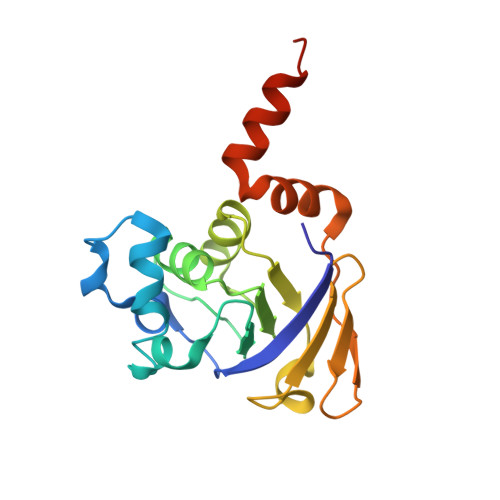
Crystal structure of LeuD from Methanococcus jannaschii.
Lee, E.H., Cho, Y.W., Hwang, K.Y.(2012) Biochem Biophys Res Commun 419: 160-164
- PubMed: 22326391 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2012.01.125
- Primary Citation Related Structures:
3VBA - PubMed Abstract:
3-Isopropylmalate/citramalate (IPM) isomerase catalyzes the second step in the leucine biosynthesis pathway. IPM isomerase from Methanococcus jannaschii is a complex protein consisting of a large (MjLeuC) and a small subunit (MjLeuD). It has broad substrate specificity, unlike other bacterial IPM isomerases. In order to understand the reasons for this broad substrate specificity, we determined the crystal structure of MjLeuD at a resolution of 2.0 Å. The asymmetric unit contained 6 molecules of LeuD, including three homodimers. The overall structure had a β/β/α sandwich-fold consisting of 8 α-helices and 7 β-strands. The C-terminal helix, which is important in homodimer formation, showed conformational differences between two homodimer forms of MjLeuD. In addition, we identified a hydrophobic residue (Val28) near the substrate recognition region that may explain the broad substrate specificity of IPM isomerase. Therefore, we suggest that LeuD proteins can be divided into 2 subfamilies, LeuD subfamilies 1 and 2, which show differences in overall structure and in the substrate recognition region.
- Division of Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 136-701, South Korea.
Organizational Affiliation: