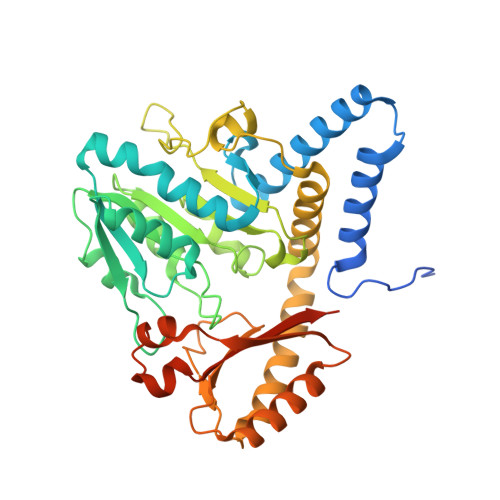
Crystal structure of the cysteine desulfurase DndA from Streptomyces lividans which is involved in DNA phosphorothioation
Chen, F., Zhang, Z., Lin, K., Qian, T., Zhang, Y., You, D., He, X., Wang, Z., Liang, J., Deng, Z., Wu, G.(2012) PLoS One 7: e36635-e36635
- PubMed: 22570733
- DOI: https://doi.org/10.1371/journal.pone.0036635
- Primary Citation of Related Structures:
3VAX - PubMed Abstract:
DNA phosphorothioation is widespread among prokaryotes, and might function to restrict gene transfer among different kinds of bacteria. There has been little investigation into the structural mechanism of the DNA phosphorothioation process. DndA is a cysteine desulfurase which is involved in the first step of DNA phosphorothioation. In this study, we determined the crystal structure of Streptomyces lividans DndA in complex with its covalently bound cofactor PLP, to a resolution of 2.4 Å. Our structure reveals the molecular mechanism that DndA employs to recognize its cofactor PLP, and suggests the potential binding site for the substrate L-cysteine on DndA. In contrast to previously determined structures of cysteine desulfurases, the catalytic cysteine of DndA was found to reside on a β strand. This catalytic cysteine is very far away from the presumable location of the substrate, suggesting that a conformational change of DndA is required during the catalysis process to bring the catalytic cysteine close to the substrate cysteine. Moreover, our in vitro enzymatic assay results suggested that this conformational change is unlikely to be a simple result of random thermal motion, since moving the catalytic cysteine two residues forward or backward in the primary sequence completely disabled the cysteine desulfurase activity of DndA.
- State Key Laboratory of Microbial Metabolism, School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, Shanghai, China.
Organizational Affiliation: