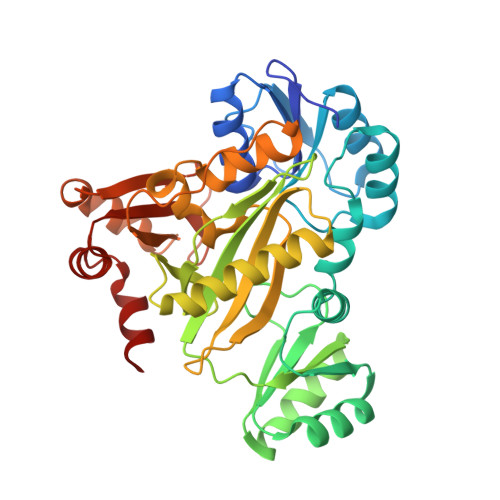
Elucidation of the bicarbonate binding site and insights into the carboxylation mechanism of (N(5))-carboxyaminoimidazole ribonucleotide synthase (PurK) from Bacillus anthracis.
Tuntland, M.L., Santarsiero, B.D., Johnson, M.E., Fung, L.W.(2014) Acta Crystallogr D Biol Crystallogr 70: 3057-3065
- PubMed: 25372694
- DOI: https://doi.org/10.1107/S1399004714021166
- Primary Citation of Related Structures:
3QFF, 3R5H, 3V4S, 4DLK - PubMed Abstract:
Structures of (N(5))-carboxyaminoimidazole ribonucleotide synthase (PurK) from Bacillus anthracis with various combinations of ATP, ADP, Mg(2+), bicarbonate and aminoimidazole ribonucleotide (AIR) in the active site are presented. The binding site of bicarbonate has only been speculated upon previously, but is shown here for the first time. The binding involves interactions with the conserved residues Arg272, His274 and Lys348. These structures provide insights into each ligand in the active site and allow a possible mechanism to be proposed for the reaction that converts bicarbonate and AIR, in the presence of ATP, to produce (N(5))-carboxyaminoimidazole ribonucleotide. The formation of a carboxyphosphate intermediate through ATP phosphoryl transfer is proposed, followed by carboxylation of AIR to give the product, facilitated by a cluster of conserved residues and an active-site water network.
- Department of Chemistry, University of Illinois at Chicago, Chicago, IL 60607, USA.
Organizational Affiliation: