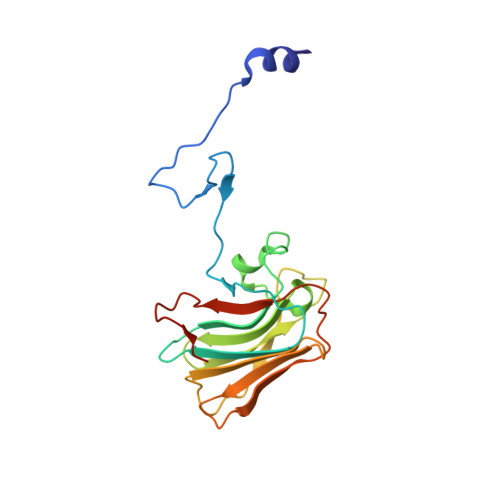
Structure of the rhesus monkey TRIM5alpha PRYSPRY domain, the HIV capsid recognition module.
Biris, N., Yang, Y., Taylor, A.B., Tomashevski, A., Guo, M., Hart, P.J., Diaz-Griffero, F., Ivanov, D.N.(2012) Proc Natl Acad Sci U S A 109: 13278-13283
- PubMed: 22847415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1203536109
- Primary Citation Related Structures:
2LM3, 3UV9 - PubMed Abstract:
Tripartite motif protein TRIM5α blocks retroviral replication after cell entry, and species-specific differences in its activity are determined by sequence variations within the C-terminal B30.2/PRYSPRY domain. Here we report a high-resolution structure of a TRIM5α PRYSPRY domain, the PRYSPRY of the rhesus monkey TRIM5α that potently restricts HIV infection, and identify features involved in its interaction with the HIV capsid. The extensive capsid-binding interface maps on the structurally divergent face of the protein formed by hypervariable loop segments, confirming that TRIM5α evolution is largely determined by its binding specificity. Interactions with the capsid are mediated by flexible variable loops via a mechanism that parallels antigen recognition by IgM antibodies, a similarity that may help explain some of the unusual functional properties of TRIM5α. Distinctive features of this pathogen-recognition interface, such as structural plasticity conferred by the mobile v1 segment and interaction with multiple epitopes, may allow restriction of divergent retroviruses and increase resistance to capsid mutations.
- Department of Biochemistry, University of Texas Health Science Center at San Antonio, San Antonio, TX 78229, USA.
Organizational Affiliation: