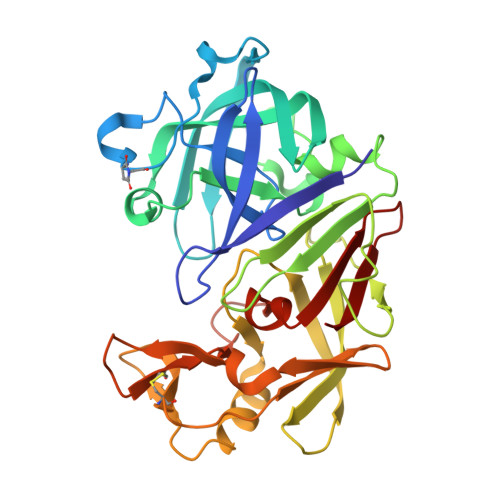
An analysis of subdomain orientation, conformational change and disorder in relation to crystal packing of aspartic proteinases.
Bailey, D., Carpenter, E.P., Coker, A., Coker, S., Read, J., Jones, A.T., Erskine, P., Aguilar, C.F., Badasso, M., Toldo, L., Rippmann, F., Sanz-Aparicio, J., Albert, A., Blundell, T.L., Roberts, N.B., Wood, S.P., Cooper, J.B.(2012) Acta Crystallogr D Biol Crystallogr 68: 541-552
- PubMed: 22525752 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912004817
- Primary Citation Related Structures:
3URI, 3URJ, 3URL, 3UTL - PubMed Abstract:
The analysis reported here describes detailed structural studies of endothiapepsin (the aspartic proteinase from Endothia parasitica), with and without bound inhibitors, and human pepsin 3b. Comparison of multiple crystal structures of members of the aspartic proteinase family has revealed small but significant differences in domain orientation in different crystal forms. In this paper, it is shown that these differences in domain orientation do not necessarily correlate with the presence or absence of bound inhibitors, but appear to stem at least partly from crystal contacts mediated by sulfate ions. However, since the same inherent flexibility of the structure is observed for other enzymes in this family such as human pepsin, the native structure of which is also reported here, the observed domain movements may well have implications for the mechanism of catalysis.
- Incisive Media, 32-34 Broadwick Street, London W1A 2HG, England.
Organizational Affiliation: