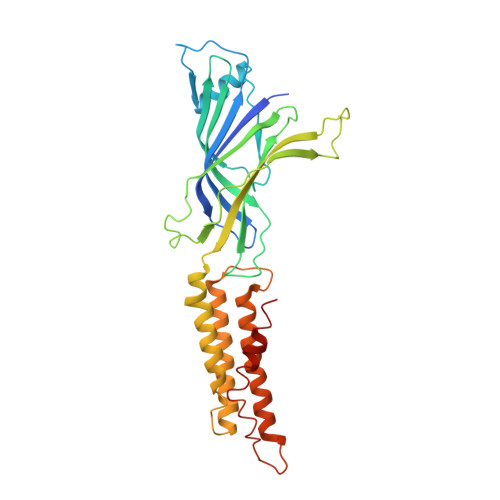
Mutations that stabilize the open state of the Erwinia chrisanthemi ligand-gated ion channel fail to change the conformation of the pore domain in crystals.
Gonzalez-Gutierrez, G., Lukk, T., Agarwal, V., Papke, D., Nair, S.K., Grosman, C.(2012) Proc Natl Acad Sci U S A 109: 6331-6336
- PubMed: 22474383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1119268109
- Primary Citation Related Structures:
3UQ4, 3UQ5, 3UQ7 - PubMed Abstract:
The determination of structural models of the various stable states of an ion channel is a key step toward the characterization of its conformational dynamics. In the case of nicotinic-type receptors, different structures have been solved but, thus far, these different models have been obtained from different members of the superfamily. In the case of the bacterial member ELIC, a cysteamine-gated channel from Erwinia chrisanthemi, a structural model of the protein in the absence of activating ligand (and thus, conceivably corresponding to the closed state of this channel) has been previously generated. In this article, electrophysiological characterization of ELIC mutants allowed us to identify pore mutations that slow down the time course of desensitization to the extent that the channel seems not to desensitize at all for the duration of the agonist applications (>20 min). Thus, it seems reasonable to conclude that the probability of ELIC occupying the closed state is much lower for the ligand-bound mutants than for the unliganded wild-type channel. To gain insight into the conformation adopted by ELIC under these conditions, we solved the crystal structures of two of these mutants in the presence of a concentration of cysteamine that elicits an intracluster open probability of >0.9. Curiously, the obtained structural models turned out to be nearly indistinguishable from the model of the wild-type channel in the absence of bound agonist. Overall, our findings bring to light the limited power of functional studies in intact membranes when it comes to inferring the functional state of a channel in a crystal, at least in the case of the nicotinic-receptor superfamily.
- Department of Molecular and Integrative Physiology, Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA.
Organizational Affiliation: