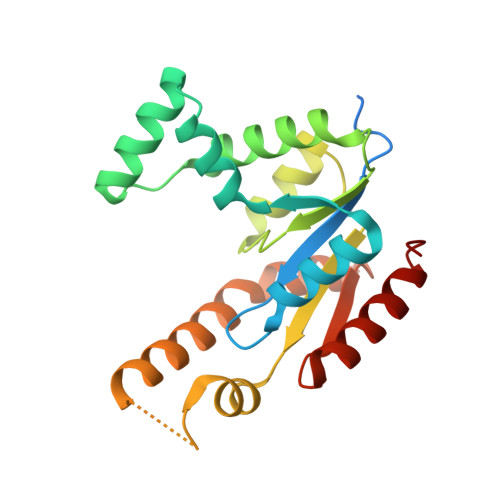
Structural and kinetic studies of Schistosoma mansoni adenylate kinases.
Marques, I., Romanello, L., Demarco, R., Pereira, H.D.(2012) Mol Biochem Parasitol 185: 157-160
- PubMed: 22841753
- DOI: https://doi.org/10.1016/j.molbiopara.2012.07.003
- Primary Citation Related Structures:
3UMF - PubMed Abstract:
The human parasite Schistosoma mansoni is totally dependent on the purine salvage pathway in order to supply large quantities of purine precursors for its energy and DNA biosynthetic needs. Adenylate kinase (ADK) is responsible for the conversion of AMP (produced by the adenosine kinase reaction) into ADP, which is subsequently converted into ATP by nucleoside diphosphate kinase (NDPK). ADK and NDPK are the most active enzymes of the pathway, probably reflecting an evolutionary adaptation due to the intense use of the branch in which they participate. However, notwithstanding their importance very little information has been accumulated found regarding these enzymes. In this work two adenylate kinases from S. mansoni were cloned and heterologously expressed in Escherichia coli. The purified products were utilized in activity assays, and displayed kinetic parameters similar to the corresponding human orthologous proteins. The cytosolic S. mansoni ADK was crystallized and its structure solved allowing us to detect a difference in the nucleotide binding site when compared with the human ortholog.
- Instituto de Física, Universidade Federal de Goiás, CP 131, 74001-970 Goiânia, GO, Brazil.
Organizational Affiliation: