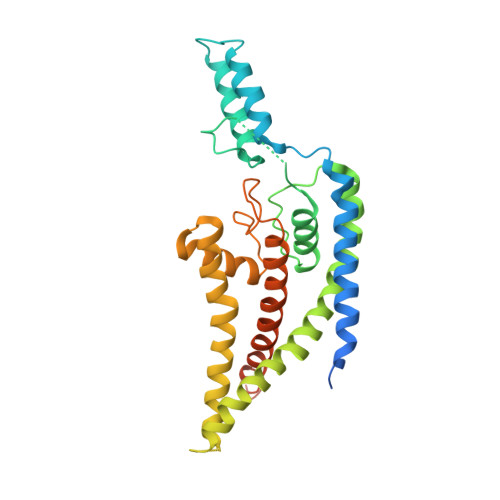
Crystal structure of the human K2P TRAAK, a lipid- and mechano-sensitive K+ ion channel.
Brohawn, S.G., del Marmol, J., MacKinnon, R.(2012) Science 335: 436-441
- PubMed: 22282805 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1213808
- Primary Citation Related Structures:
3UM7 - PubMed Abstract:
TRAAK channels, members of the two-pore domain K(+) (potassium ion) channel family K2P, are expressed almost exclusively in the nervous system and control the resting membrane potential. Their gating is sensitive to polyunsaturated fatty acids, mechanical deformation of the membrane, and temperature changes. Physiologically, these channels appear to control the noxious input threshold for temperature and pressure sensitivity in dorsal root ganglia neurons. We present the crystal structure of human TRAAK at a resolution of 3.8 angstroms. The channel comprises two protomers, each containing two distinct pore domains, which create a two-fold symmetric K(+) channel. The extracellular surface features a helical cap, 35 angstroms tall, that creates a bifurcated pore entryway and accounts for the insensitivity of two-pore domain K(+) channels to inhibitory toxins. Two diagonally opposed gate-forming inner helices form membrane-interacting structures that may underlie this channel's sensitivity to chemical and mechanical properties of the cell membrane.
- Laboratory of Molecular Neurobiology and Biophysics and Howard Hughes Medical Institute, The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: