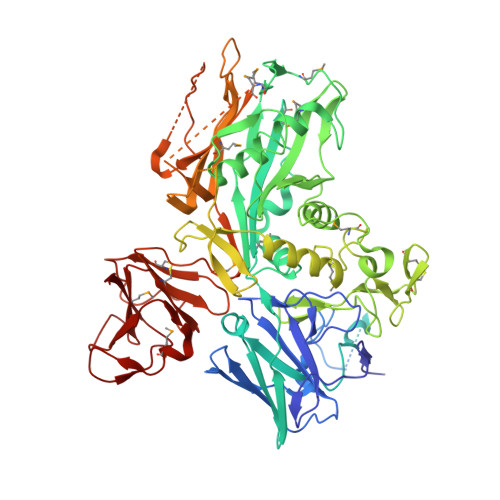
Structural Insight into the Bacterial Mucinase StcE Essential to Adhesion and Immune Evasion during Enterohemorrhagic E. coli Infection.
Yu, A.C., Worrall, L.J., Strynadka, N.C.(2012) Structure 20: 707-717
- PubMed: 22483117
- DOI: https://doi.org/10.1016/j.str.2012.02.015
- Primary Citation Related Structures:
3UJZ, 4DNY - PubMed Abstract:
Mucin glycoproteins with large numbers of O-linked glycosylations comprise the mucosal barrier lining the mammalian gastrointestinal tract from mouth to gut. A critical biological function of mucins is to protect the underlying epithelium from infection. Enterohemorrhagic Escherichia coli (EHEC), the mediator of severe food- and water-borne disease, can breach this barrier and adhere to intestinal cells. StcE, a ∼100 kDa metalloprotease secreted by EHEC, plays a pivotal role in remodeling the mucosal lining during infection. To obtain mechanistic insight into its function, we have determined the structure of StcE. Our data reveal a dynamic, multidomain architecture featuring an unusually large substrate-binding cleft and a prominent polarized surface charge distribution highly suggestive of an electrostatic role in substrate targeting. The observation of key conserved motifs in the active site allows us to propose the structural basis for the specific recognition of α-O-glycan-containing substrates. Complementary biochemical analysis provides further insight into its distinct substrate specificity and binding stoichiometry.
- Department of Biochemistry and Molecular Biology and the Center for Blood Research, Life Sciences Center, 2350 Health Sciences Mall, University of British Columbia, Vancouver, BC V6T 1Z3, Canada.
Organizational Affiliation: