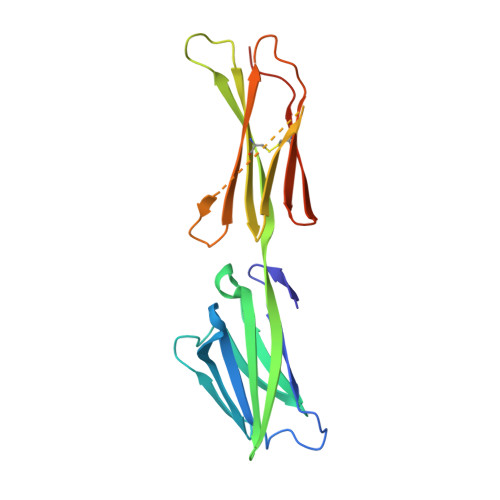
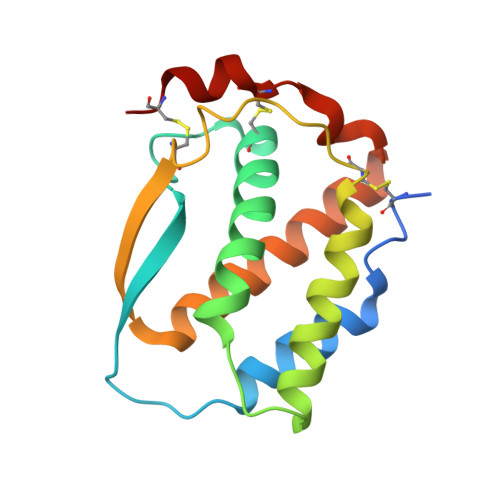
Allosteric competitive inactivation of hematopoietic CSF-1 signaling by the viral decoy receptor BARF1
Elegheert, J., Bracke, N., Pouliot, P., Gutsche, I., Shkumatov, A.V., Tarbouriech, N., Verstraete, K., Bekaert, A., Burmeister, W.P., Svergun, D.I., Lambrecht, B.N., Vergauwen, B., Savvides, S.N.(2012) Nat Struct Mol Biol 19: 938-947
- PubMed: 22902366 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2367
- Primary Citation Related Structures:
3UEZ, 3UF2, 3UF5, 4ADF, 4ADQ - PubMed Abstract:
Hematopoietic human colony-stimulating factor 1 (hCSF-1) is essential for innate and adaptive immunity against viral and microbial infections and cancer. The human pathogen Epstein-Barr virus secretes the lytic-cycle protein BARF1 that neutralizes hCSF-1 to achieve immunomodulation. Here we show that BARF1 binds the dimer interface of hCSF-1 with picomolar affinity, away from the cognate receptor-binding site, to establish a long-lived complex featuring three hCSF-1 at the periphery of the BARF1 toroid. BARF1 locks dimeric hCSF-1 into an inactive conformation, rendering it unable to signal via its cognate receptor on human monocytes. This reveals a new functional role for hCSF-1 cooperativity in signaling. We propose a new viral strategy paradigm featuring an allosteric decoy receptor of the competitive type, which couples efficient sequestration and inactivation of the host growth factor to abrogate cooperative assembly of the cognate signaling complex.
- Laboratory for Protein Biochemistry and Biomolecular Engineering, Department of Biochemistry & Microbiology, Ghent University, Ghent, Belgium.
Organizational Affiliation: