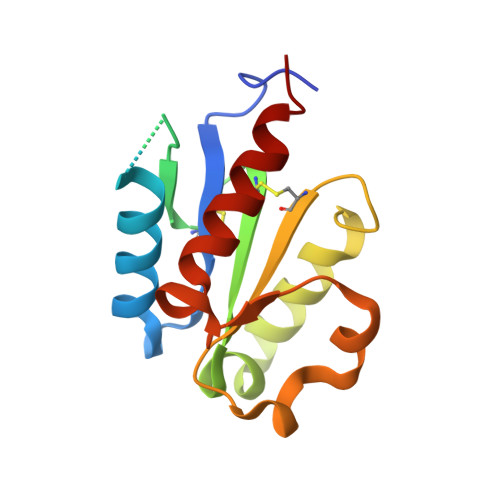
Structural Insights into TIR Domain Specificity of the Bridging Adaptor Mal in TLR4 Signaling
Lin, Z., Lu, J., Zhou, W., Shen, Y.(2012) PLoS One 7: e34202-e34202
- PubMed: 22485159
- DOI: https://doi.org/10.1371/journal.pone.0034202
- Primary Citation Related Structures:
3UB2, 3UB3, 3UB4 - PubMed Abstract:
MyD88 adaptor-like protein (Mal) is a crucial adaptor that acts as a bridge to recruit the MyD88 molecule to activated TLR4 receptors in response to invading pathogens. The specific assembly of the Toll/interleukin-1 receptor (TIR) domains of TLR4, Mal and MyD88 is responsible for proper signal transduction in the TLR4 signaling pathway. However, the molecular mechanism for the specificity of these TIR domains remains unclear. Here, we present the crystal structure of the TIR domain of the human Mal molecule (Mal-TIR) at a resolution of 2.4 Å. Unexpectedly, Mal-TIR exhibits an extraordinarily long AB loop, but no αB helix or BB loop, distinguishing it from other TIR domains. More importantly, the Mal-TIR AB loop is capable of mediating direct binding to the TIR domains of TLR4 and MyD88 simultaneously. We also found that Mal-TIR can form a back-to-back dimer that may resemble the dimeric assembly of the entire Mal molecule. Our data demonstrate the bridge role of the Mal-TIR domain and provide important information about TIR domain specificity.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin, China.
Organizational Affiliation: