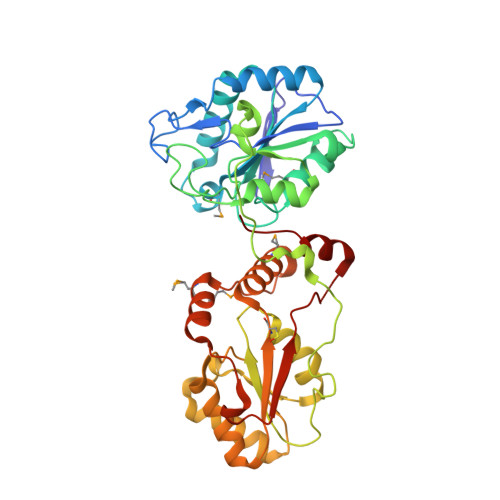
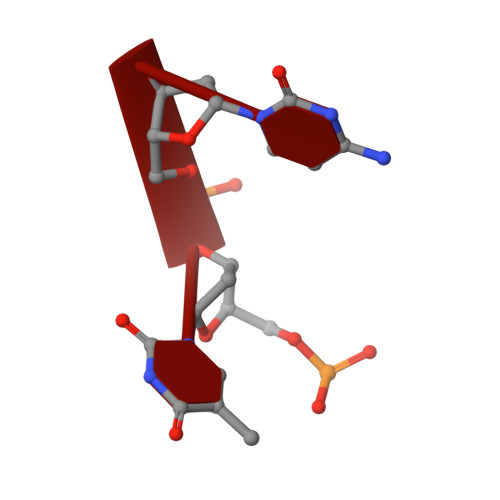
Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Coquelle, N., Havali-Shahriari, Z., Bernstein, N., Green, R., Glover, J.N.(2011) Proc Natl Acad Sci U S A 108: 21022-21027
- PubMed: 22171004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1112036108
- Primary Citation Related Structures:
3U7E, 3U7F, 3U7G, 3U7H - PubMed Abstract:
Polynucleotide kinase/phosphatase (PNKP) is a critical mammalian DNA repair enzyme that generates 5'-phosphate and 3'-hydroxyl groups at damaged DNA termini that are required for subsequent processing by DNA ligases and polymerases. The PNKP phosphatase domain recognizes 3'-phosphate termini within DNA nicks, gaps, or at double- or single-strand breaks. Here we present a mechanistic rationale for the recognition of damaged DNA termini by the PNKP phosphatase domain. The crystal structures of PNKP bound to single-stranded DNA substrates reveals a narrow active site cleft that accommodates a single-stranded substrate in a sequence-independent manner. Biochemical studies suggest that the terminal base pairs of double-stranded substrates near the 3'-phosphate are destabilized by PNKP to allow substrate access to the active site. A positively charged surface distinct from the active site specifically facilitates interactions with double-stranded substrates, providing a complex DNA binding surface that enables the recognition of diverse substrates.
- Department of Biochemistry, University of Alberta, Edmonton, AB, Canada T6G 2H7.
Organizational Affiliation: