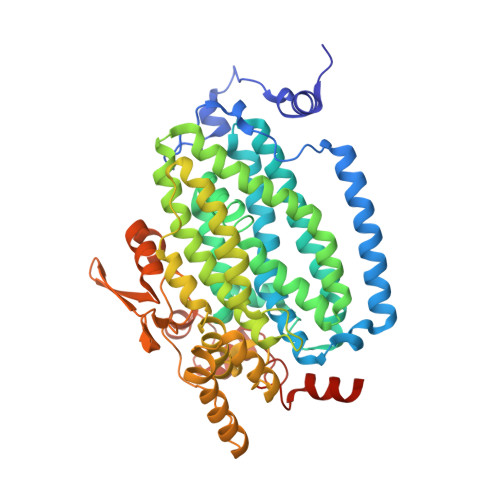
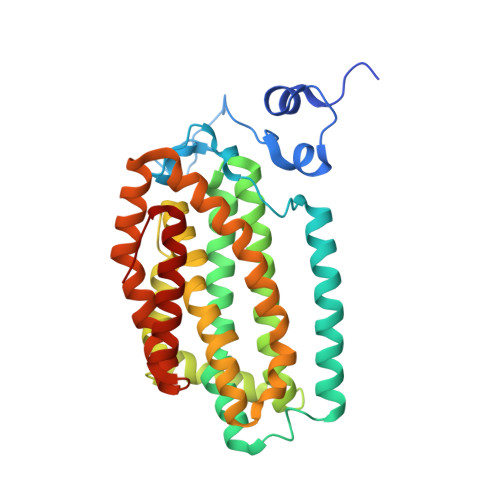
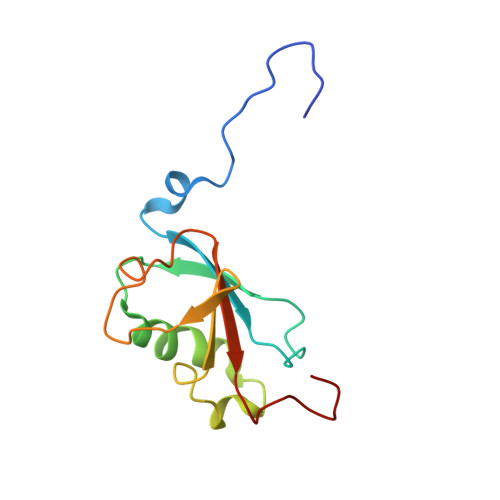
Analysis of Substrate Access to Active Sites in Bacterial Multicomponent Monooxygenase Hydroxylases: X-ray Crystal Structure of Xenon-Pressurized Phenol Hydroxylase from Pseudomonas sp. OX1.
McCormick, M.S., Lippard, S.J.(2011) Biochemistry 50: 11058-11069
- PubMed: 22136180 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi201248b
- Primary Citation Related Structures:
3U52 - PubMed Abstract:
In all structurally characterized bacterial multicomponent monooxygenase (BMM) hydroxylase proteins, a series of hydrophobic cavities in the α-subunit trace a conserved path from the protein exterior to the carboxylate-bridged diiron active site. This study examines these cavities as a potential route for transport of dioxygen to the active site by crystallographic characterization of a xenon-pressurized sample of the hydroxylase component of phenol hydroxylase from Pseudomonas sp. OX1. Computational analyses of the hydrophobic cavities in the hydroxylase α-subunits of phenol hydroxylase (PHH), soluble methane monooxygenase (MMOH), and toluene/o-xylene monooxygenase (ToMOH) are also presented. The results, together with previous findings from crystallographic studies of xenon-pressurized sMMO hydroxylase, clearly identify the propensity for these cavities to bind hydrophobic gas molecules in the protein interior. This proposed functional role is supported by recent stopped flow kinetic studies of ToMOH variants [Song, W. J., et al. (2011) Proc. Natl. Acad. Sci. U.S.A.108, 14795-14800]. In addition to information about the Xe sites, the structure determination revealed significantly weakened binding of regulatory protein to the hydroxylase in comparison to that in the previously reported structure of PHH, as well as the presence of a newly identified metal-binding site in the α-subunit that adopts a linear coordination environment consistent with Cu(I), and a glycerol molecule bound to Fe1 in a fashion that is unique among hydrocarbon-diiron site adducts reported to date in BMM hydroxylase structures. Finally, a comparative analysis of the α-subunit structures of PHH, MMOH, and ToMOH details proposed routes for the other three BMM substrates, the hydrocarbon, electrons, and protons, comprising cavities, channels, hydrogen-bonding networks, and pores in the structures of their α-subunits.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States.
Organizational Affiliation: