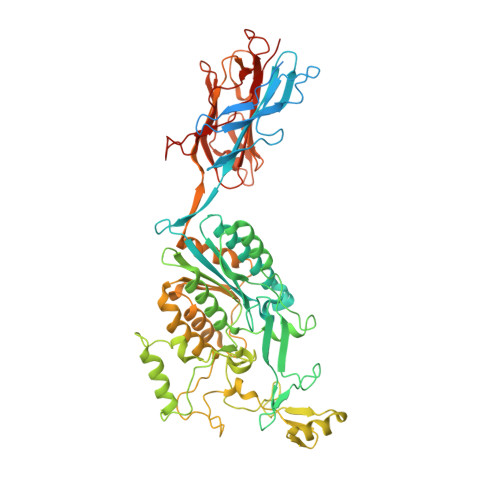
Structure of Streptococcus agalactiae tip pilin GBS104: a model for GBS pili assembly and host interactions.
Krishnan, V., Dwivedi, P., Kim, B.J., Samal, A., Macon, K., Ma, X., Mishra, A., Doran, K.S., Ton-That, H., Narayana, S.V.(2013) Acta Crystallogr D Biol Crystallogr 69: 1073-1089
- PubMed: 23695252 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913004642
- Primary Citation Related Structures:
3TVY, 3TW0, 3TXA - PubMed Abstract:
The crystal structure of a 75 kDa central fragment of GBS104, a tip pilin from the 2063V/R strain of Streptococcus agalactiae (group B streptococcus; GBS), is reported. In addition, a homology model of the remaining two domains of GBS104 was built and a model of full-length GBS104 was generated by combining the homology model (the N1 and N4 domains) and the crystal structure of the 75 kDa fragment (the N2 and N3 domains). This rod-shaped GBS104 model is constructed of three IgG-like domains (the N1, N2 and N4 domains) and one vWFA-like domain (the N3 domain). The N1 and N2 domains of GBS104 are assembled with distinct and remote segments contributed by the N- and C-termini. The metal-binding site in the N3 domain of GBS104 is in the closed/low-affinity conformation. Interestingly, this domain hosts two long arms that project away from the metal-binding site. Using site-directed mutagenesis, two cysteine residues that lock the N3 domain of GBS104 into the open/high-affinity conformation were introduced. Both wild-type and disulfide-locked recombinant proteins were tested for binding to extracellular matrix proteins such as collagen, fibronectin, fibrinogen and laminin, and an increase in fibronectin binding affinity was identified for the disulfide-locked N3 domain, suggesting that induced conformational changes may play a possible role in receptor binding.
- UNESCO Regional Centre for Biotechnology (RCB), Gurgaon 122 016, Haryana, India.
Organizational Affiliation: