Glycolipids that Elicit IFN-gama-Biased Responses from Natural Killer T Cells
Tyznik, A.J., Farber, E., Girardi, E., Birkholz, A., Li, Y., Chitale, S., So, R., Arora, P., Khurana, A., Wang, J., Porcelli, S.A., Zajonc, D.M., Kronenberg, M., Howell, A.R.(2011) Chem Biol 18: 1620-1630
- PubMed: 22195564 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2011.10.015
- Primary Citation Related Structures:
3TVM - PubMed Abstract:
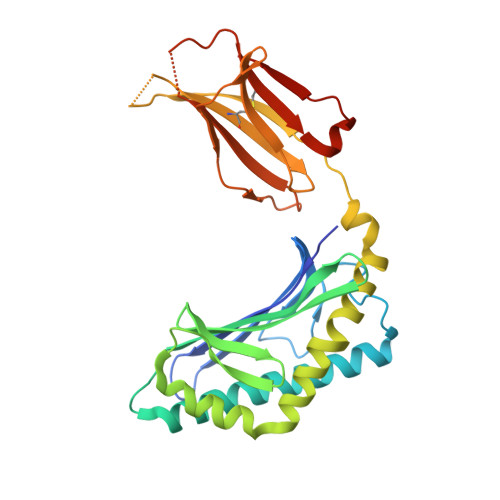
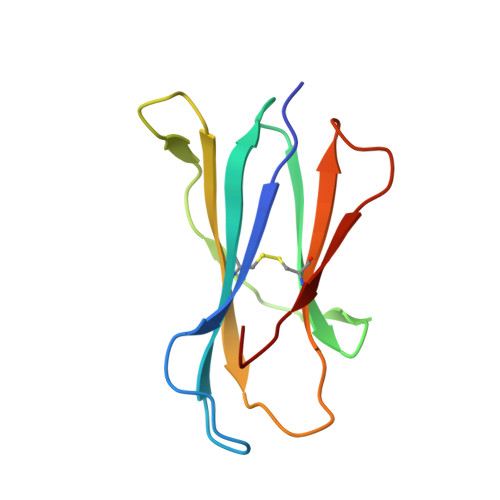
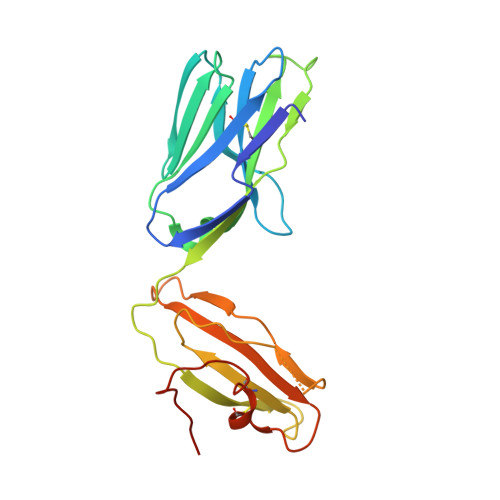
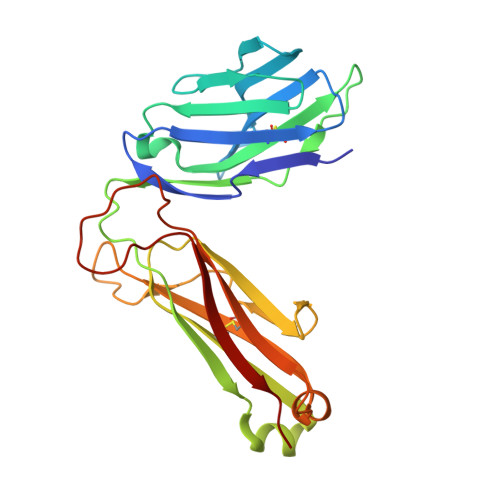
Natural killer T (NKT) cells recognize glycolipids presented by CD1d. The first antigen described, α-galactosyl ceramide (αGalCer), is a potential anticancer agent whose activity depends upon IFN-γ secretion. We report two analogs of αGalCer based on a naturally occurring glycosphingolipid, plakoside A. These compounds induce enhanced IFN-γ that correlates with detergent-resistant binding to CD1d and an increased stability of the lipid-CD1d complexes on antigen-presenting cells. Structural analysis on one of the analogs indicates that it is more deeply bound inside the CD1d groove, suggesting tighter lipid-CD1d interactions. To our knowledge, this is the first example in which structural information provides an explanation for the increased lipid-CD1d stability, likely responsible for the Th1 bias. We provide insights into the mechanism of IFN-γ-inducing compounds, and because our compounds activate human NKT cells, they could have therapeutic utility.
- Division of Developmental Immunology, La Jolla Institute for Allergy & Immunology, La Jolla, CA 92037, USA.
Organizational Affiliation: