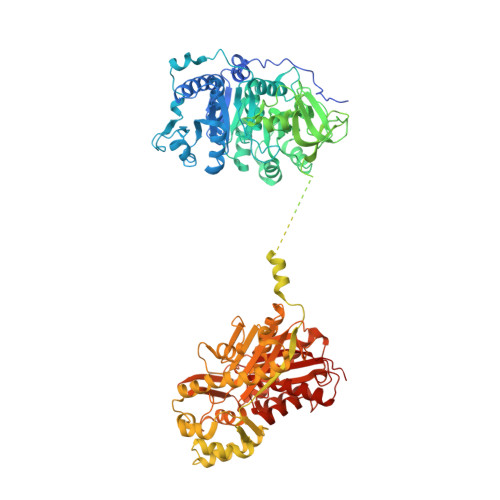
Structural and Kinetic Analysis of the Unnatural Fusion Protein 4-Coumaroyl-CoA Ligase::Stilbene Synthase.
Wang, Y., Yi, H., Wang, M., Yu, O., Jez, J.M.(2011) J Am Chem Soc 133: 20684-20687
- PubMed: 22129213
- DOI: https://doi.org/10.1021/ja2085993
- Primary Citation Related Structures:
3TSY - PubMed Abstract:
To increase the biochemical efficiency of biosynthetic systems, metabolic engineers have explored different approaches for organizing enzymes, including the generation of unnatural fusion proteins. Previous work aimed at improving the biosynthesis of resveratrol, a stilbene associated a range of health-promoting activities, in yeast used an unnatural engineered fusion protein of Arabidopsis thaliana (thale cress) 4-coumaroyl-CoA ligase (At4CL1) and Vitis vinifera (grape) stilbene synthase (VvSTS) to increase resveratrol levels 15-fold relative to yeast expressing the individual enzymes. Here we present the crystallographic and biochemical analysis of the 4CL::STS fusion protein. Determination of the X-ray crystal structure of 4CL::STS provides the first molecular view of an artificial didomain adenylation/ketosynthase fusion protein. Comparison of the steady-state kinetic properties of At4CL1, VvSTS, and 4CL::STS demonstrates that the fusion protein improves catalytic efficiency of either reaction less than 3-fold. Structural and kinetic analysis suggests that colocalization of the two enzyme active sites within 70 Å of each other provides the basis for enhanced in vivo synthesis of resveratrol.
- Donald Danforth Plant Science Center, 975 North Warson Road, St. Louis, Missouri 63132, USA.
Organizational Affiliation: